# Util libs
library(assertthat)
library(ggplot2)
library(zeallot)
library(conflicted)
library(Matrix)
# Data processing libs
if (!suppressWarnings(require(COTAN))) {
devtools::load_all("~/dev/COTAN/COTAN/")
}
conflicts_prefer(zeallot::`%->%`, zeallot::`%<-%`)
conflicts_prefer(base::setdiff)
options(parallelly.fork.enable = TRUE)
setLoggingLevel(2L)Run40: dataset cleaning and c3 clusters’ checks
Preamble
List input files
inDir <- file.path("Data/Brown_PBMC_Datasets/")
outDir <- file.path("Data/Brown_PBMC_Datasets/")
list.files(path = inDir, pattern = "\\.RDS$") [1] "capillary_blood_samples_pbmcs-Run_28.RDS"
[2] "capillary_blood_samples_pbmcs-Run_40-Cleaned.RDS"
[3] "capillary_blood_samples_pbmcs-Run_40.RDS"
[4] "capillary_blood_samples_pbmcs-Run_41-Cleaned.RDS"
[5] "capillary_blood_samples_pbmcs-Run_41.RDS"
[6] "capillary_blood_samples_pbmcs-Run_62.RDS"
[7] "capillary_blood_samples_pbmcs-Run_77-Cleaned.RDS"
[8] "capillary_blood_samples_pbmcs-Run_77.RDS"
[9] "capillary_blood_samples_pbmcs.RDS"
[10] "CD4Tcells-Run_40-Cleaned.RDS"
[11] "CD8Tcells-Run_40-Cleaned.RDS"
[12] "Run_40Cl_0CD4TcellsRawData.RDS"
[13] "Run_40Cl_1CD4TcellsRawData.RDS"
[14] "Run_40Cl_5CD4TcellsRawData.RDS"
[15] "Run_40Cl_7CD4TcellsRawData.RDS"
[16] "Run_40Cl_8CD4TcellsRawData.RDS"
[17] "Run_40Cl_9CD4TcellsRawData.RDS" globalCondition <- "capillary_blood_samples_pbmcs"
thisRun <- "40"
fileNameIn <- paste0(globalCondition, "-Run_", thisRun, ".RDS")
setLoggingFile(file.path(outDir, paste0("DatasetCleaning_Run", thisRun, ".log")))Load dataset
aRunObj <- readRDS(file = file.path(inDir, fileNameIn))
sampleCond <- getMetadataElement(aRunObj, tag = datasetTags()[["cond"]])[[1L]]
aRunObj <- initializeMetaDataset(aRunObj,
GEO = "Zenodo 8020792",
sequencingMethod = "10x V3.1",
sampleCondition = sampleCond)
getAllConditions(aRunObj)[1] "Sample.IDs" "Participant.IDs"
[3] "Cell.Barcoding.Runs" "Lane"
[5] "cell_barcoding_protocol" "run_lane_batch"
[7] "original_sample_id" getClusterizations(aRunObj)[1] "cell_type_level_1" "cell_type_level_2" "cell_type_level_3"
[4] "cell_type_level_4" "c1" "c2"
[7] "c3" "c4" dim(getRawData(aRunObj))[1] 36601 9025Cleaning
clean() using standard thresholds
c4Clusters <- getClusters(aRunObj,"c4")
cellsToDrop <- names(c4Clusters)[is.na(c4Clusters)]
aRunObj <- dropGenesCells(aRunObj, cells = cellsToDrop)
aRunObj <- clean(aRunObj)Check the initial plots
cellSizePlot(aRunObj, condName = "Participant.IDs")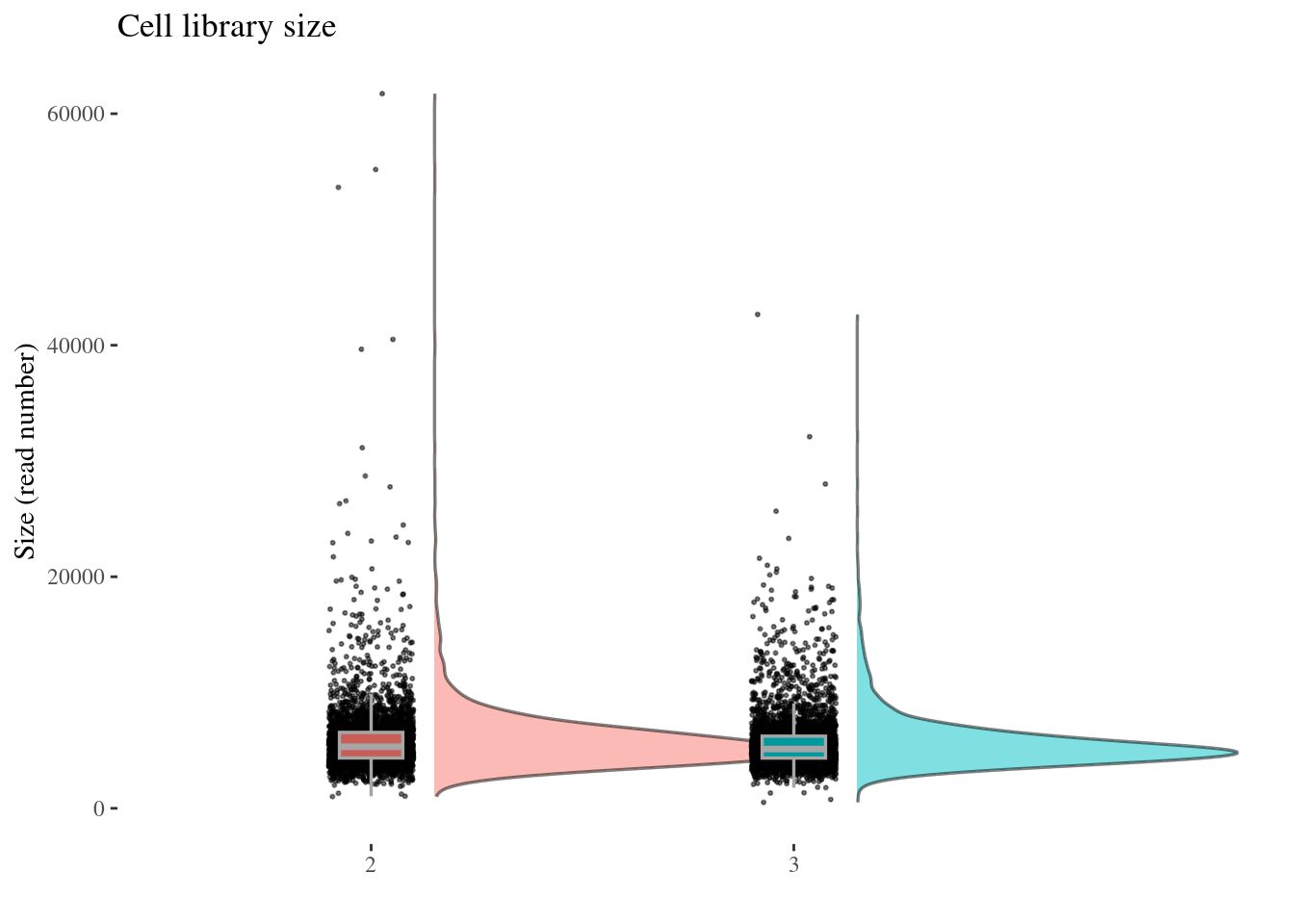
genesSizePlot(aRunObj, condName = "Participant.IDs")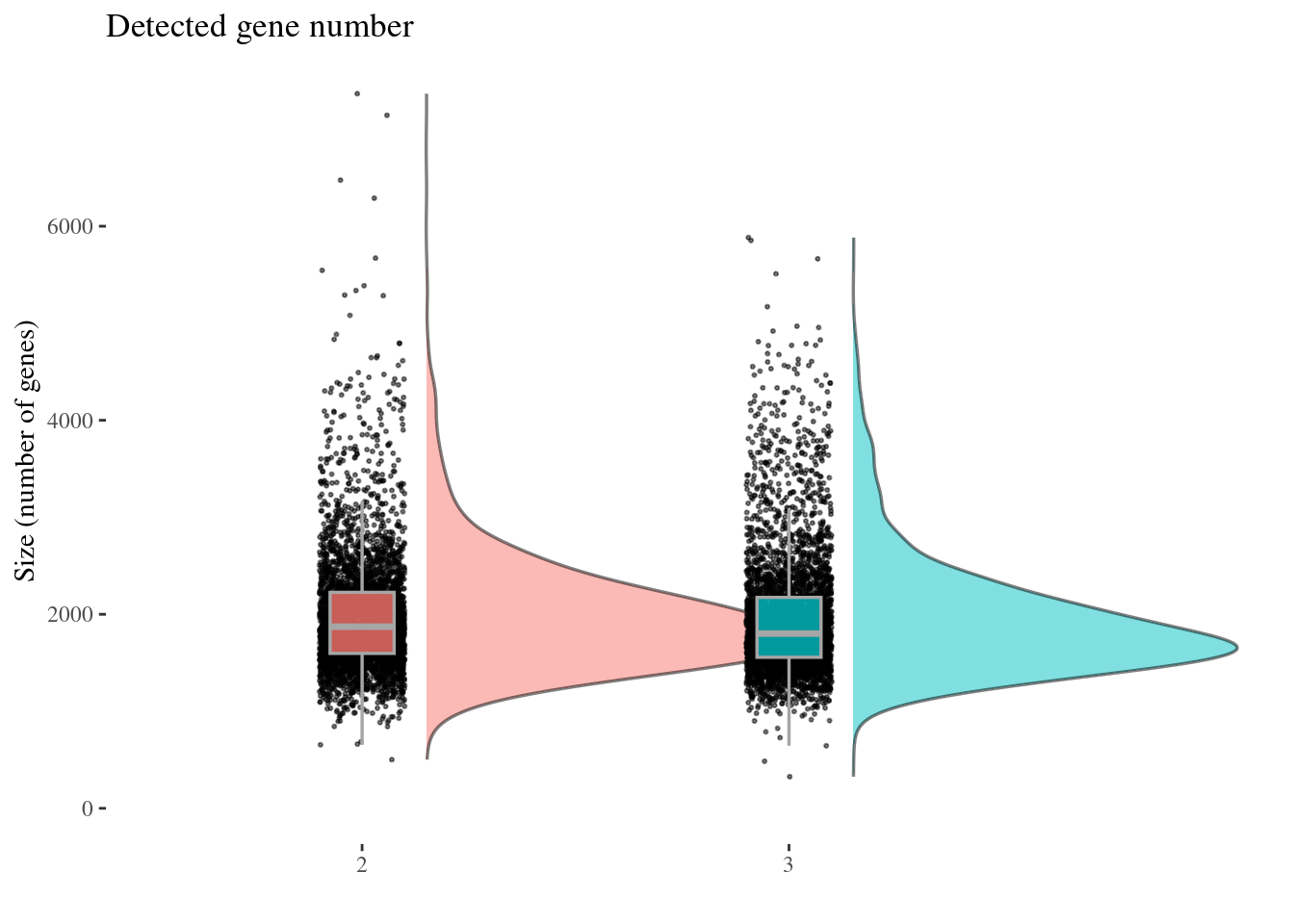
scatterPlot(aRunObj, condName = "Participant.IDs")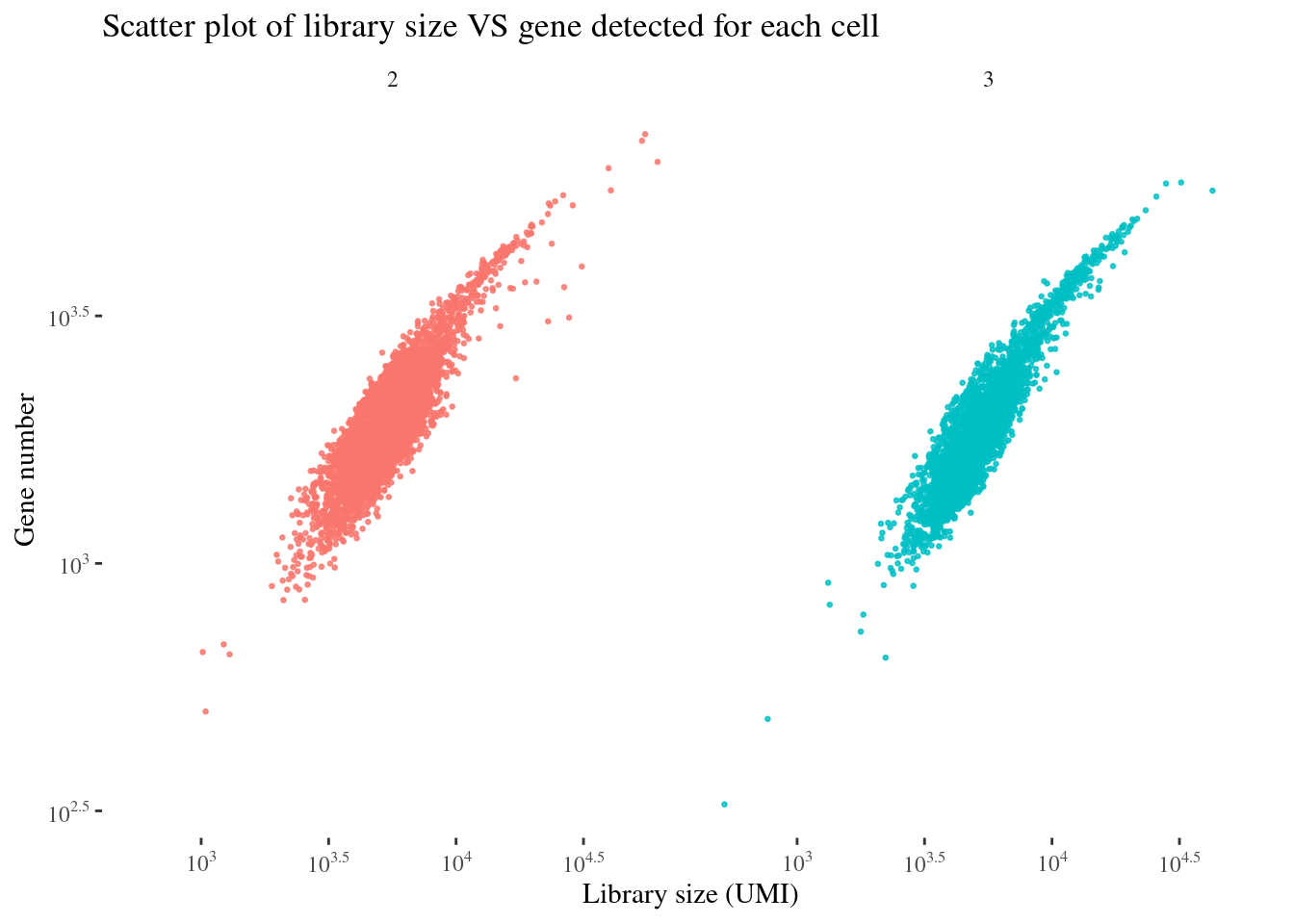
c(mitPerPlot, mitPerDf) %<-%
mitochondrialPercentagePlot(aRunObj, condName = "Participant.IDs",
genePrefix = "^MT-")
mitPerPlot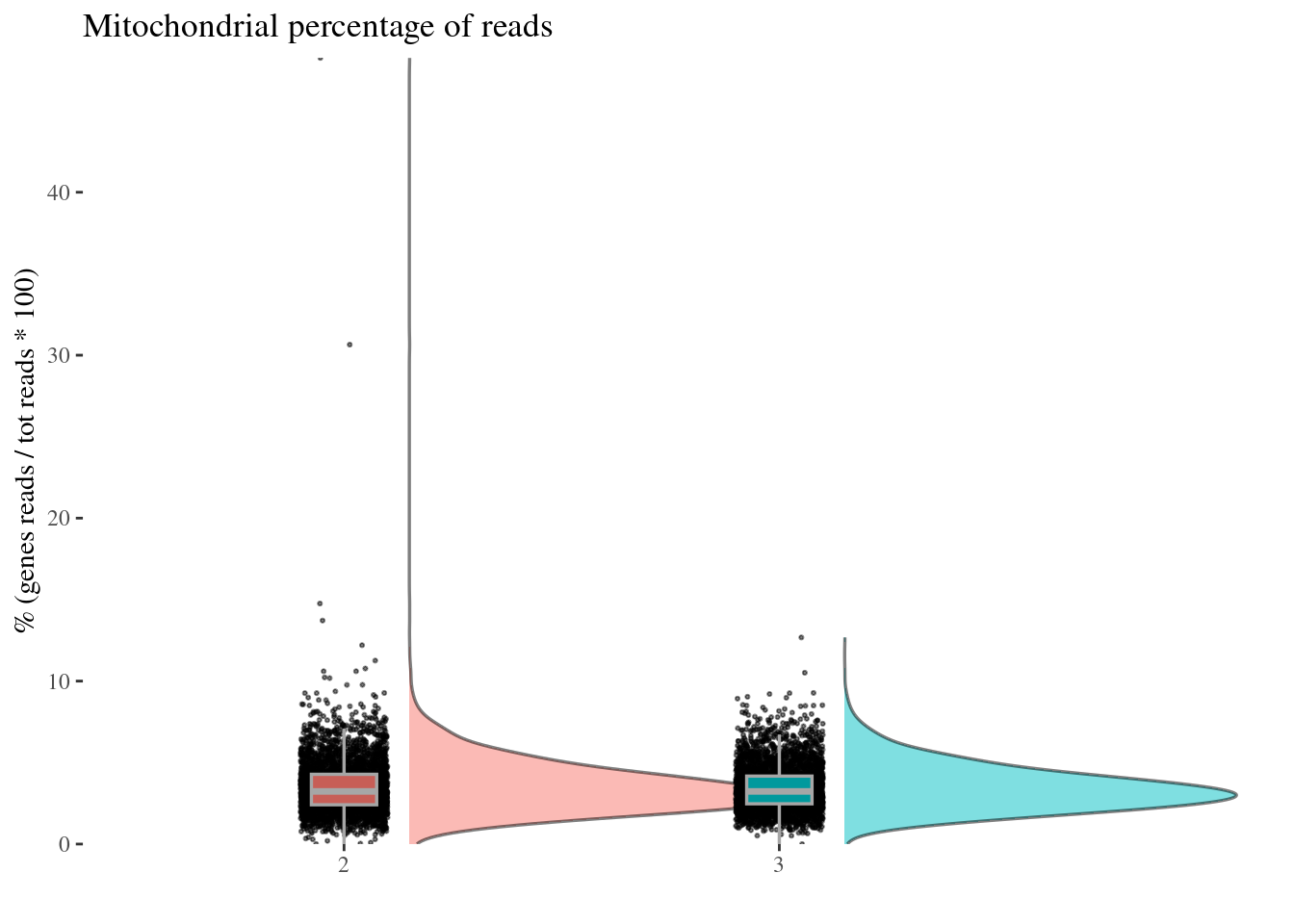
Remove too high mitochondrial percentage cells
Expected percentages are around 6%-10%: too high percentage imply dead cells
sum(mitPerDf[["mit.percentage"]] > 13.0)[1] 4sum(mitPerDf[["mit.percentage"]] > 11.0)[1] 7sum(mitPerDf[["mit.percentage"]] > 10.0)[1] 13sum(mitPerDf[["mit.percentage"]] > 8.0)[1] 64sum(mitPerDf[["mit.percentage"]] > 7.0)[1] 184mitPerThr <- 8.0
aRunObj <-
addElementToMetaDataset(
aRunObj,
"Mitochondrial percentage threshold", mitPerThr)
cellsToDrop <- getCells(aRunObj)[which(mitPerDf[["mit.percentage"]] > mitPerThr)]
aRunObj <- dropGenesCells(aRunObj, cells = cellsToDrop)Redo the plots
cellSizePlot(aRunObj, condName = "Participant.IDs")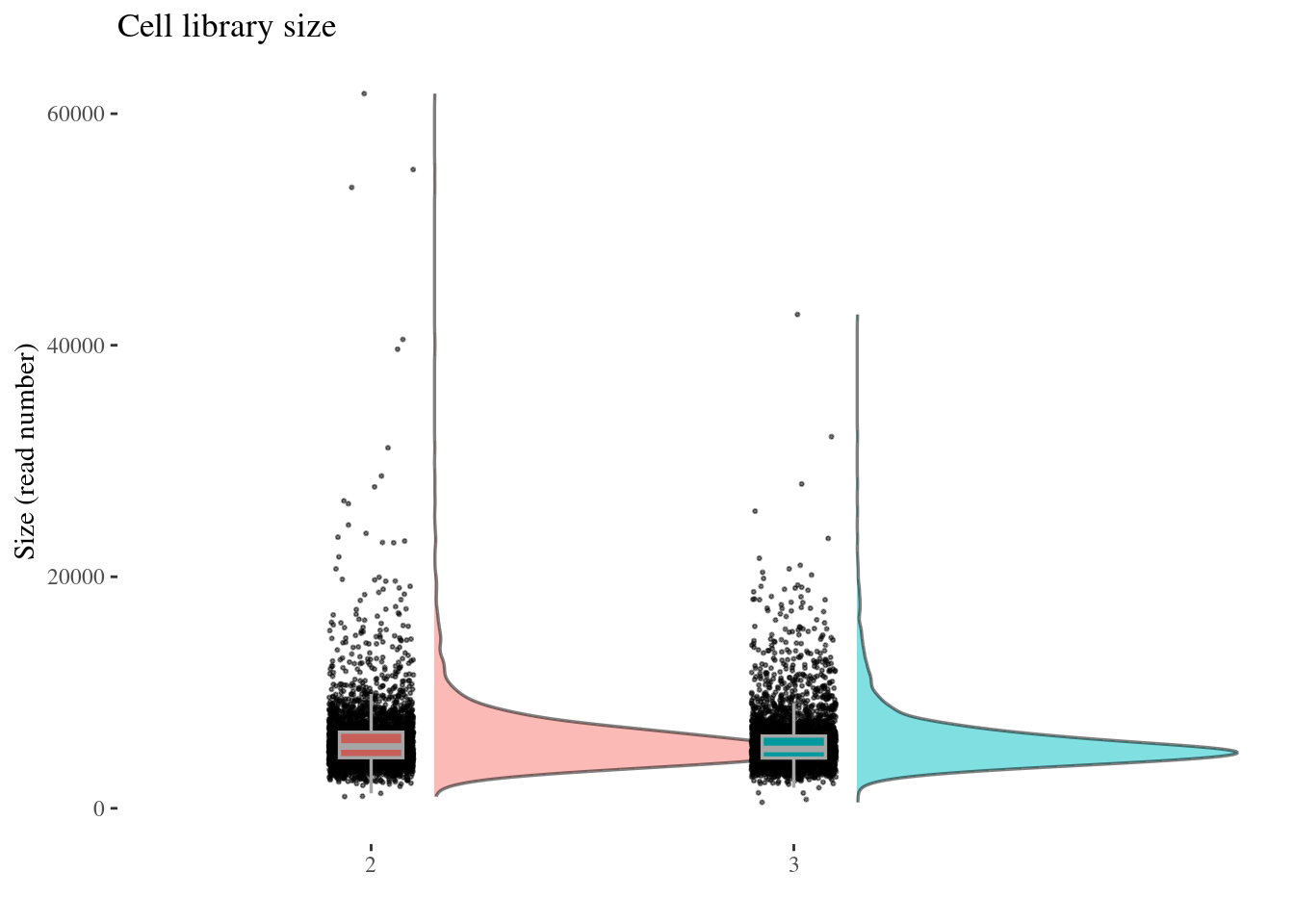
genesSizePlot(aRunObj, condName = "Participant.IDs")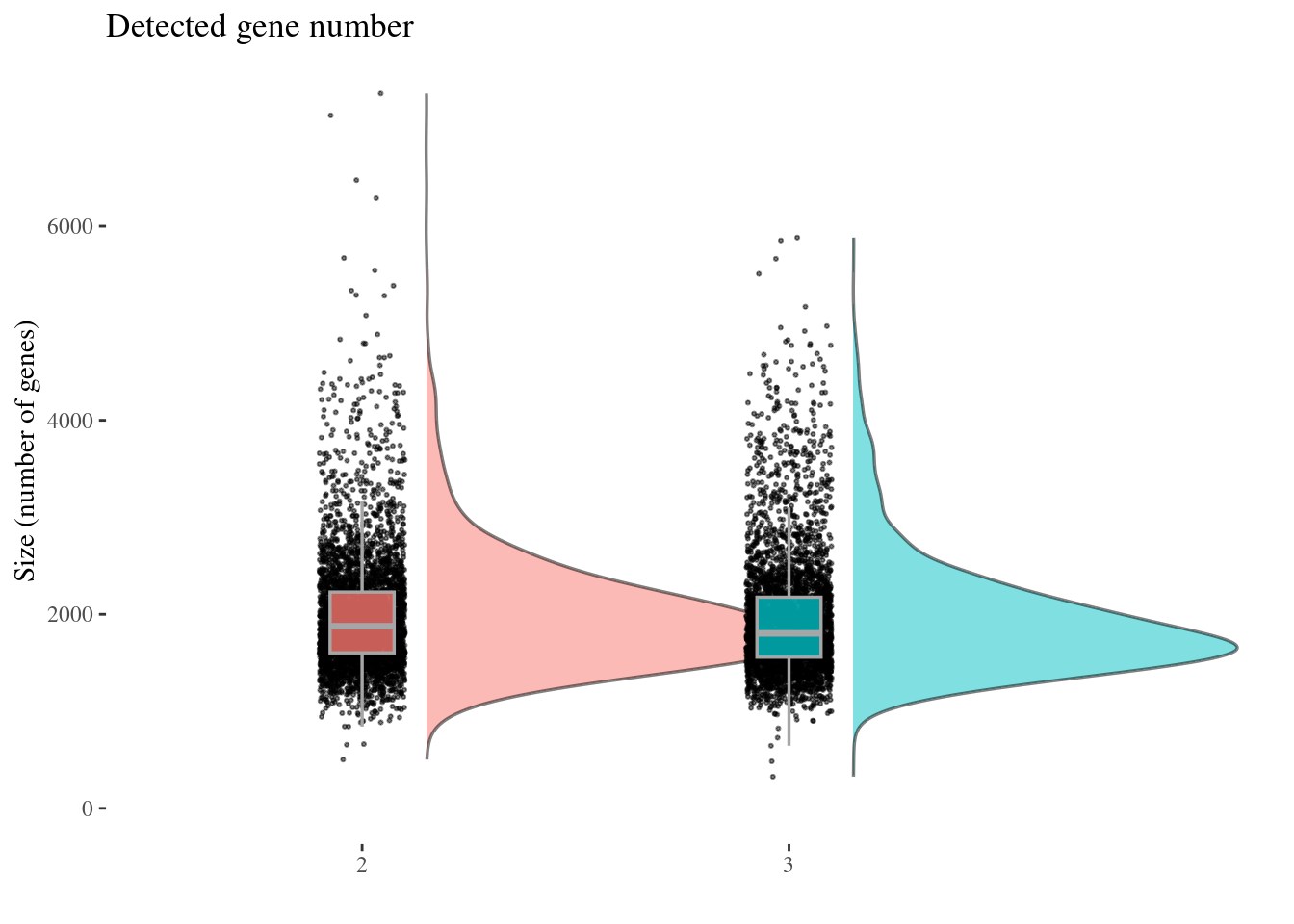
scatterPlot(aRunObj, condName = "Participant.IDs")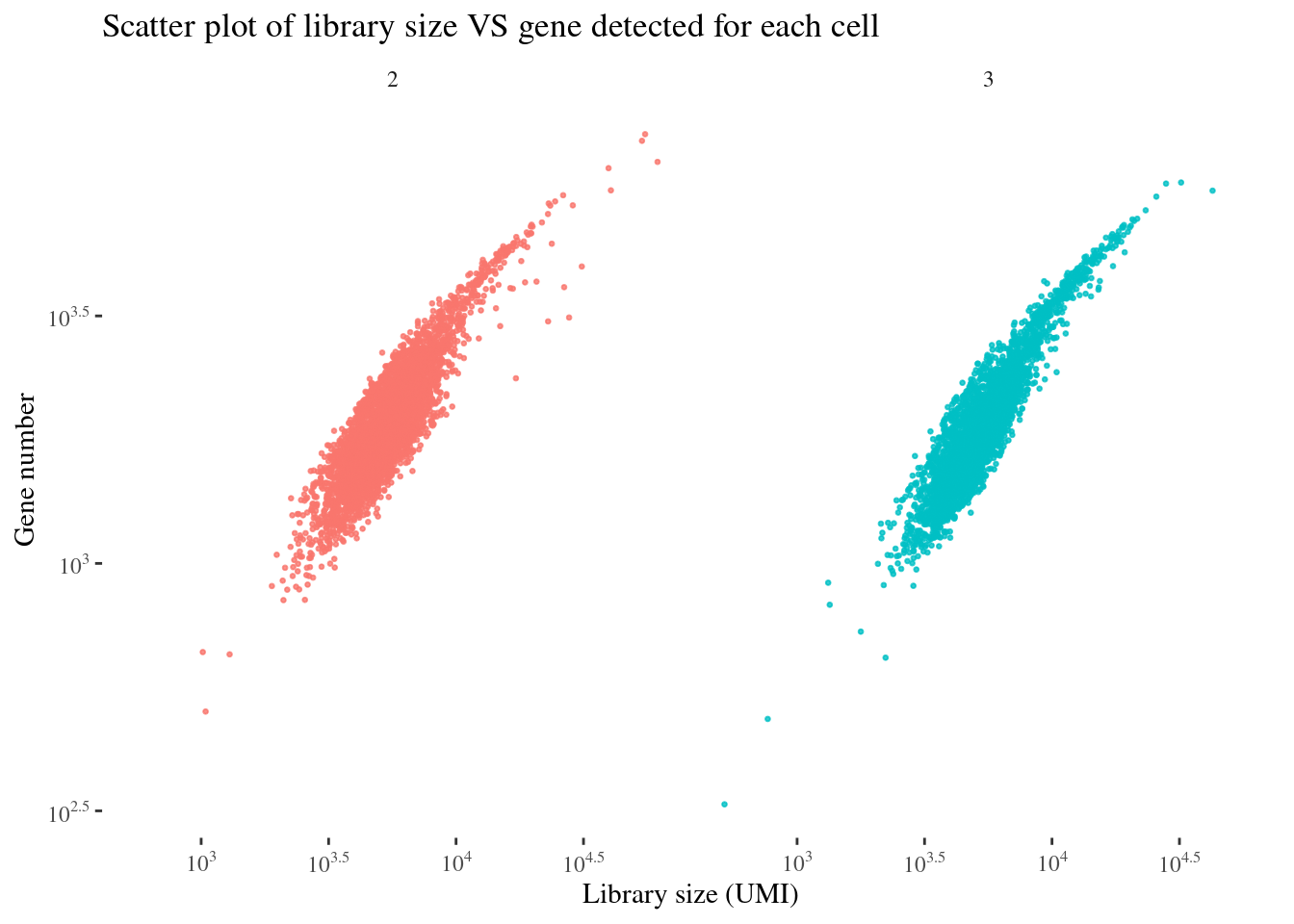
c(mitPerPlot, mitPerDf) %<-%
mitochondrialPercentagePlot(aRunObj, condName = "Participant.IDs",
genePrefix = "^MT-")
mitPerPlot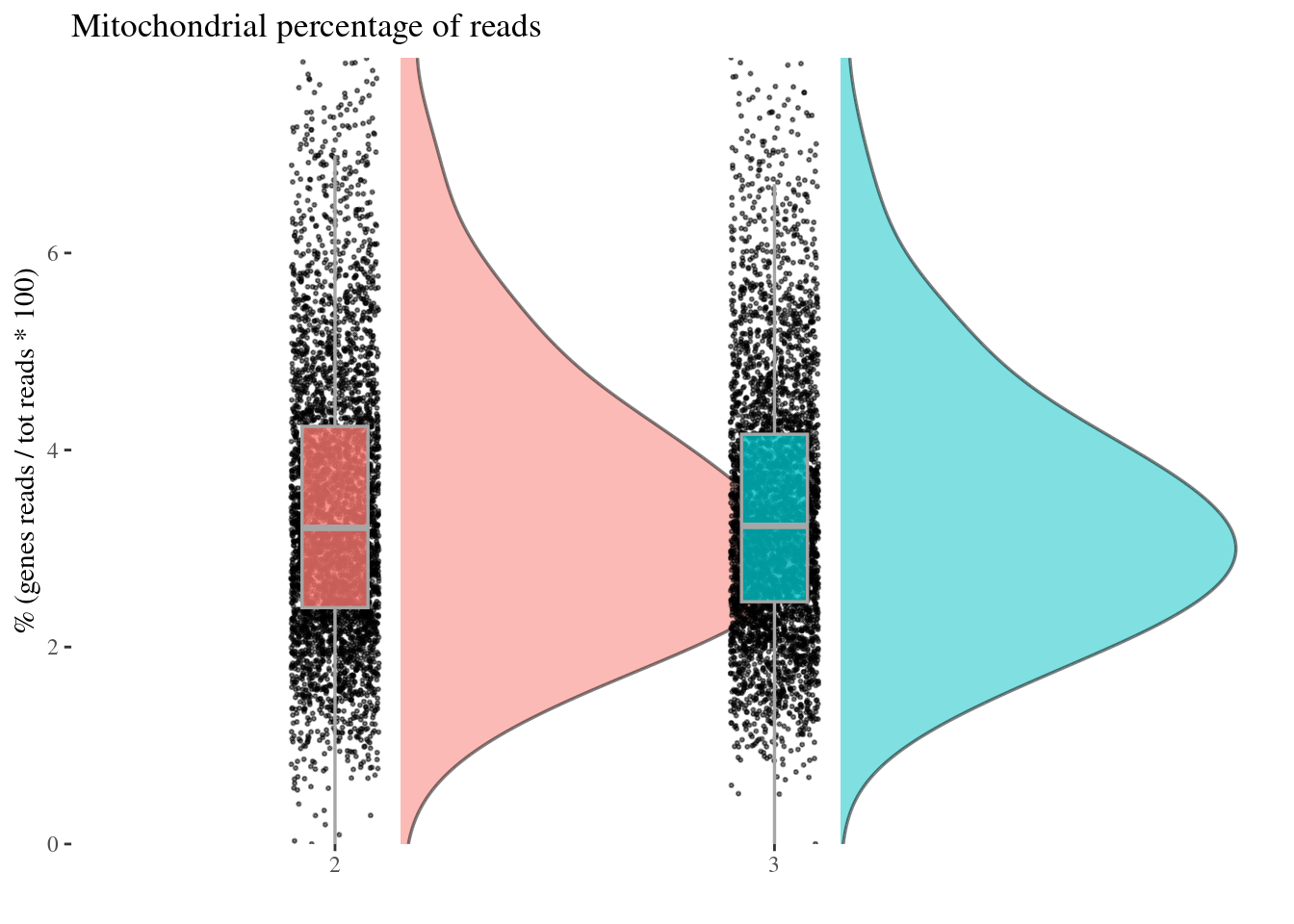
Remove cells with too high size
librarySize <- getCellsSize(aRunObj)
sum(librarySize > 25000)[1] 14sum(librarySize > 20000)[1] 28sum(librarySize > 18000)[1] 54sum(librarySize > 15000)[1] 106sum(librarySize > 10000)[1] 397cellSizeThr <- 15000
aRunObj <-
addElementToMetaDataset(
aRunObj,
"Cell size threshold", cellSizeThr)
cellsToDrop <- getCells(aRunObj)[which(librarySize > cellSizeThr)]
aRunObj <- dropGenesCells(aRunObj, cells = cellsToDrop)Redo the plots
cellSizePlot(aRunObj, condName = "Participant.IDs")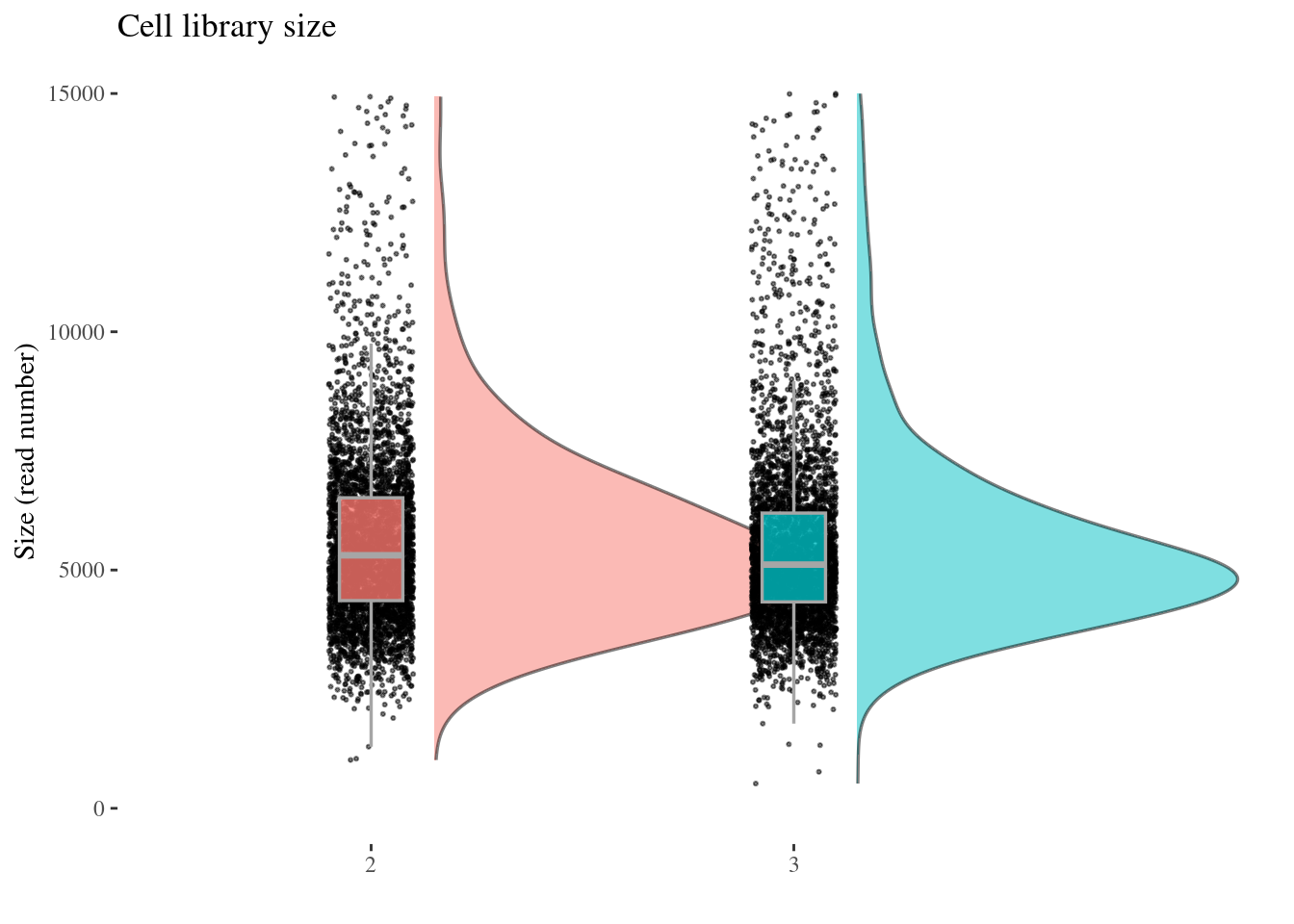
genesSizePlot(aRunObj, condName = "Participant.IDs")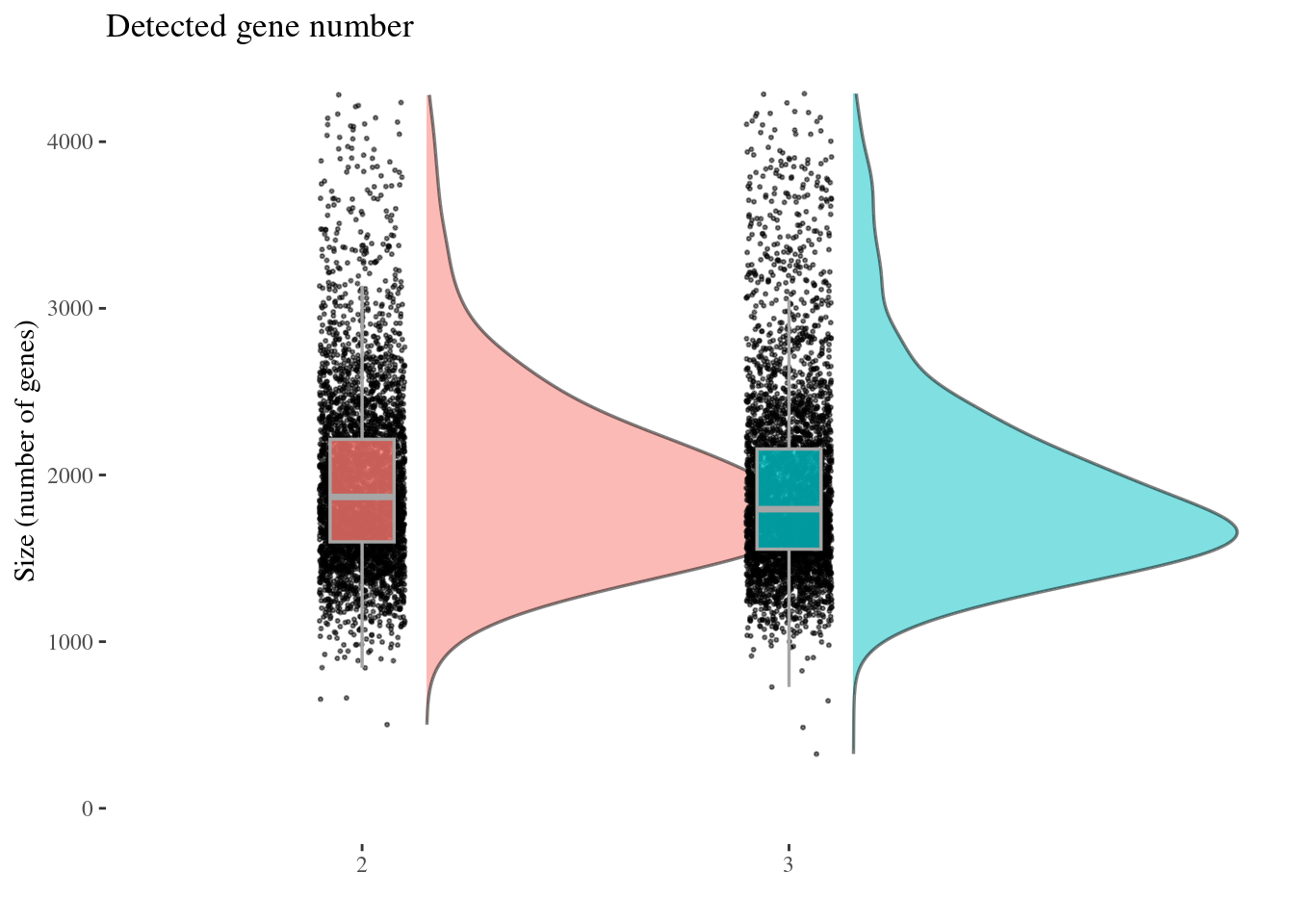
scatterPlot(aRunObj, condName = "Participant.IDs")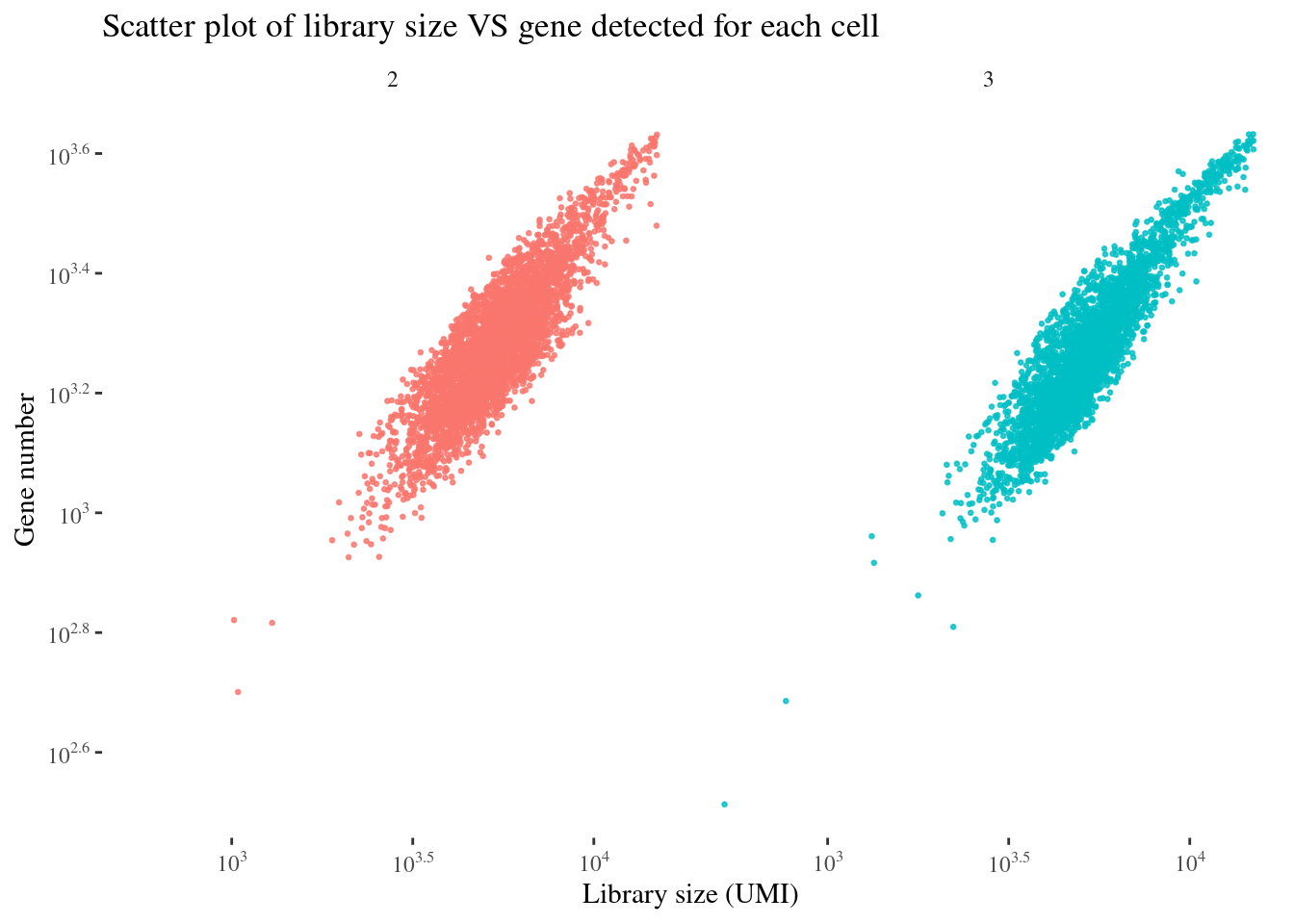
c(mitPerPlot, mitPerDf) %<-%
mitochondrialPercentagePlot(aRunObj, condName = "Participant.IDs",
genePrefix = "^MT-")
mitPerPlot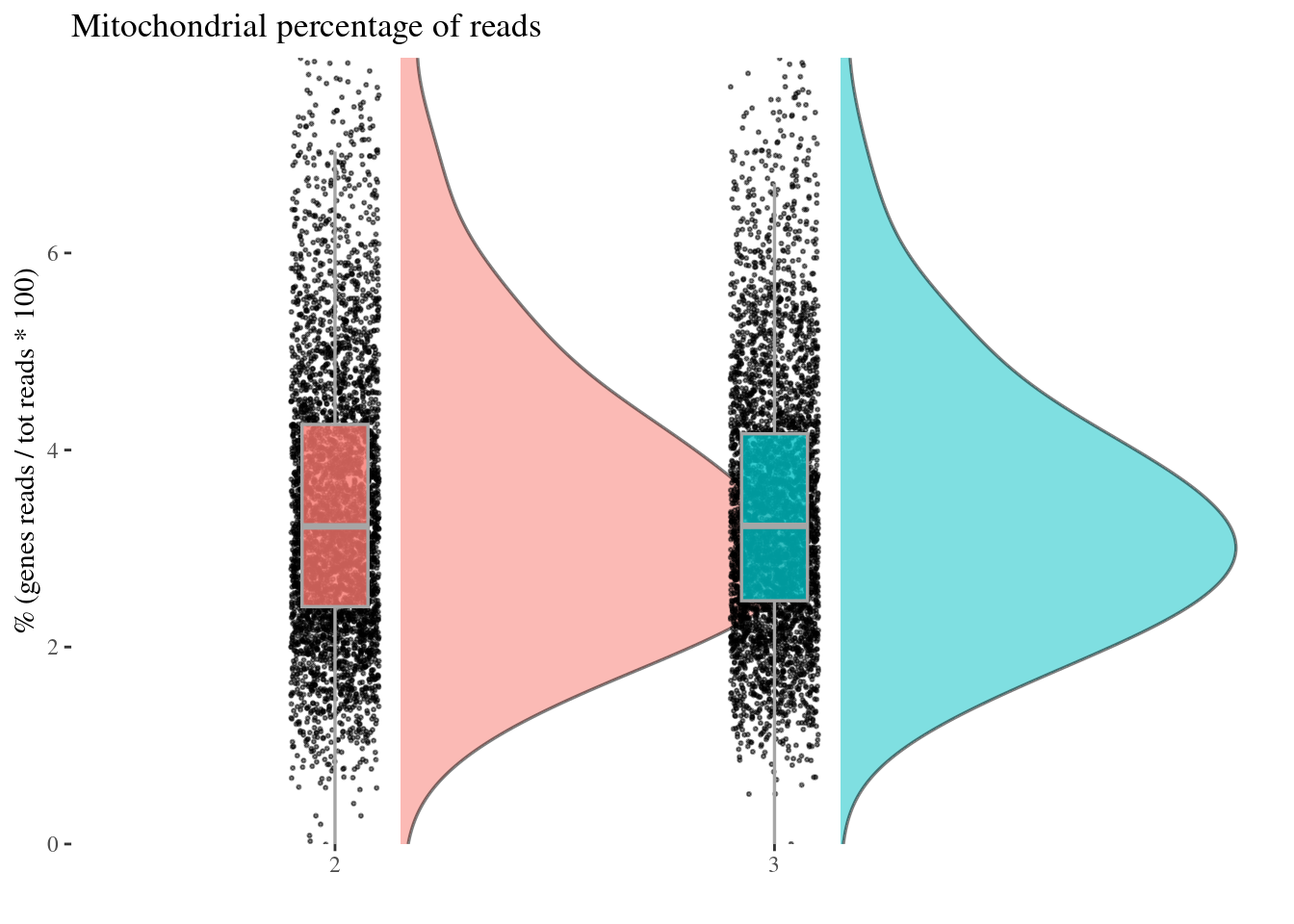
Remove cells with too low number of expressed genes
numExprGenes <- getNumExpressedGenes(aRunObj)
sum(numExprGenes < 1000)[1] 38sum(numExprGenes < 900)[1] 13sum(numExprGenes < 800)[1] 7sum(numExprGenes < 700)[1] 6sum(numExprGenes < 500)[1] 2numExprGenesThr <- 800
aRunObj <-
addElementToMetaDataset(
aRunObj,
"Num Expr Genes low threshold", numExprGenesThr)
cellsToDrop <- getCells(aRunObj)[which(numExprGenes < numExprGenesThr)]
aRunObj <- dropGenesCells(aRunObj, cells = cellsToDrop)Redo the plots
cellSizePlot(aRunObj, condName = "Participant.IDs")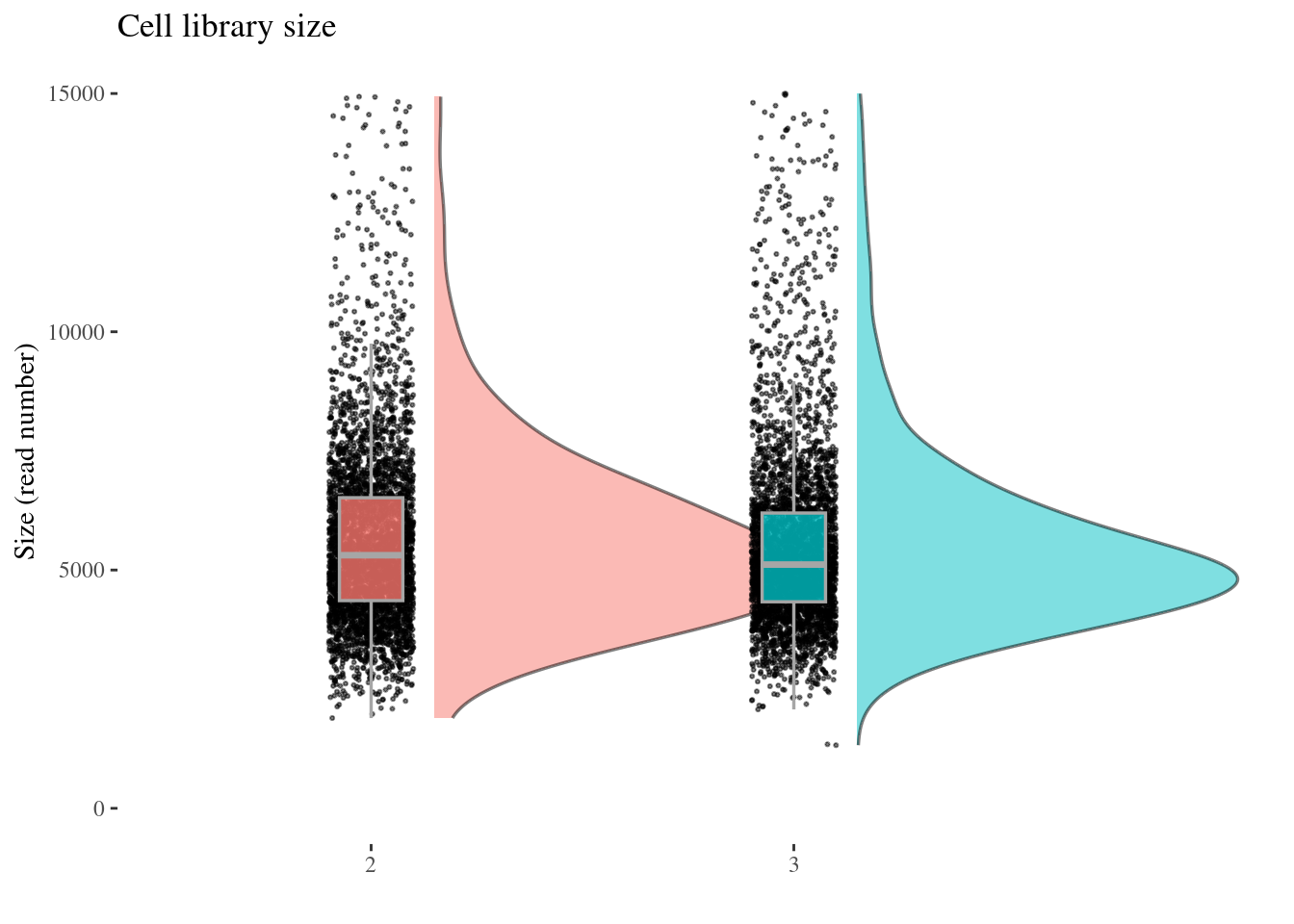
genesSizePlot(aRunObj, condName = "Participant.IDs")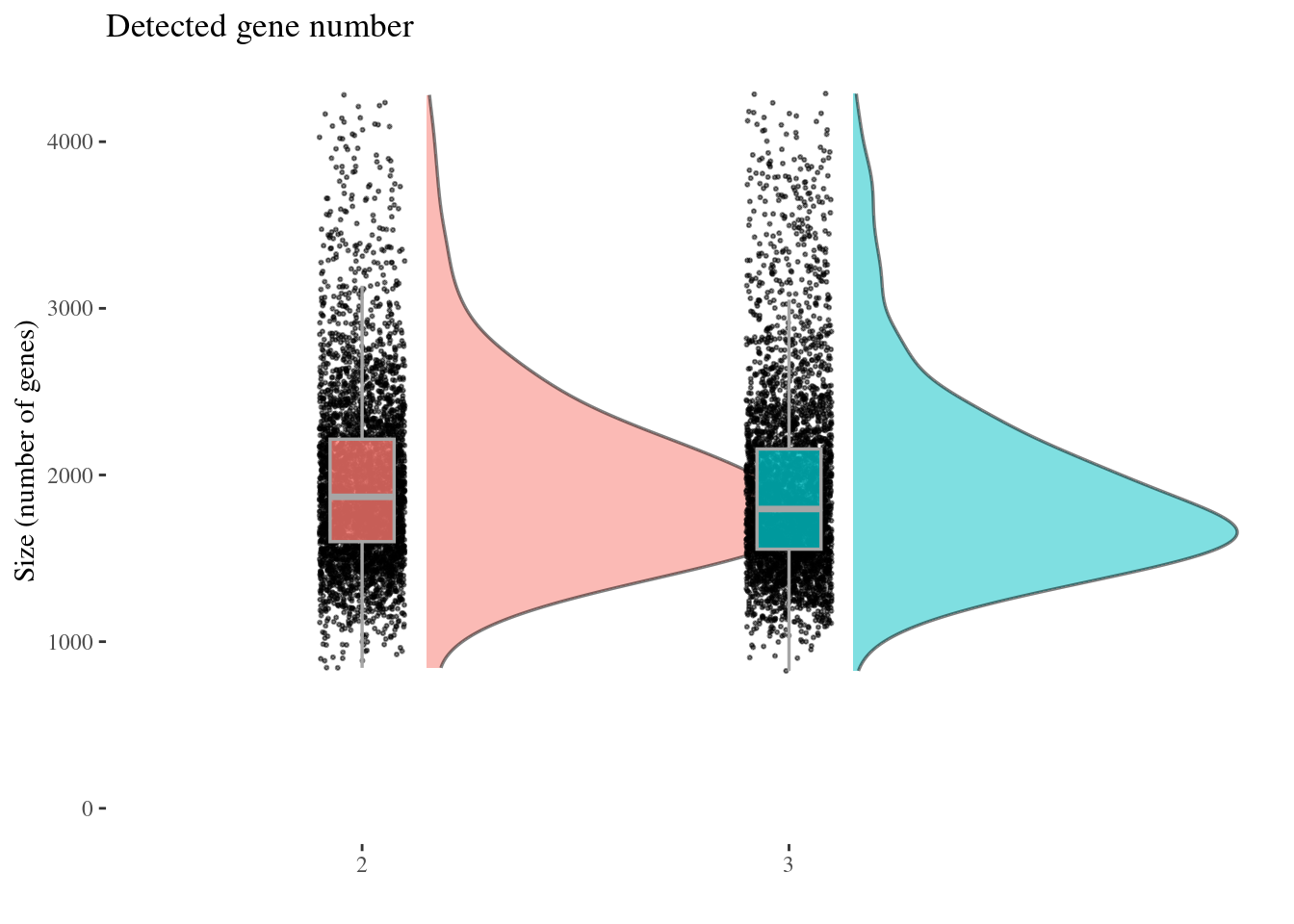
scatterPlot(aRunObj, condName = "Participant.IDs")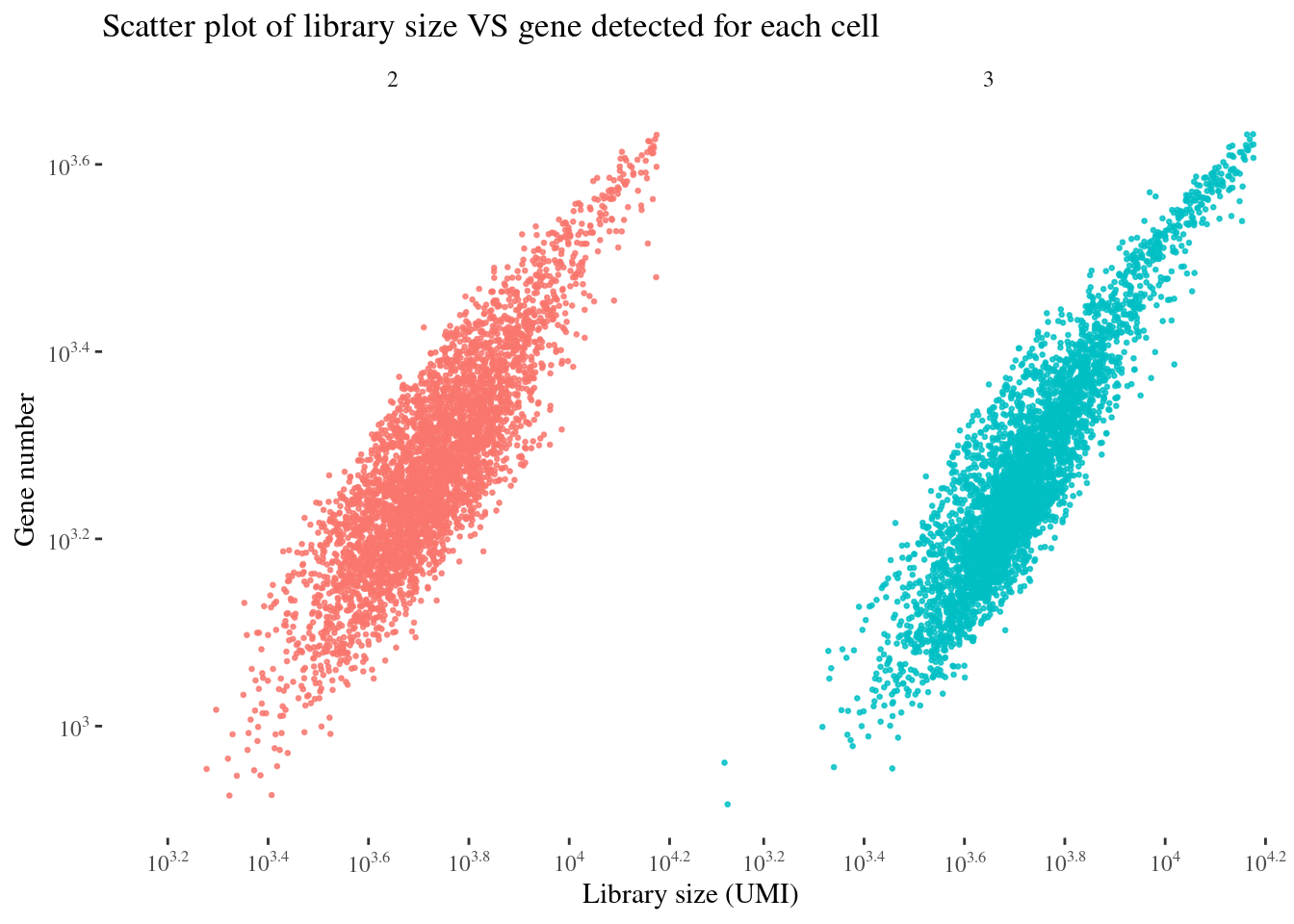
c(mitPerPlot, mitPerDf) %<-%
mitochondrialPercentagePlot(aRunObj, condName = "Participant.IDs",
genePrefix = "^MT-")
mitPerPlot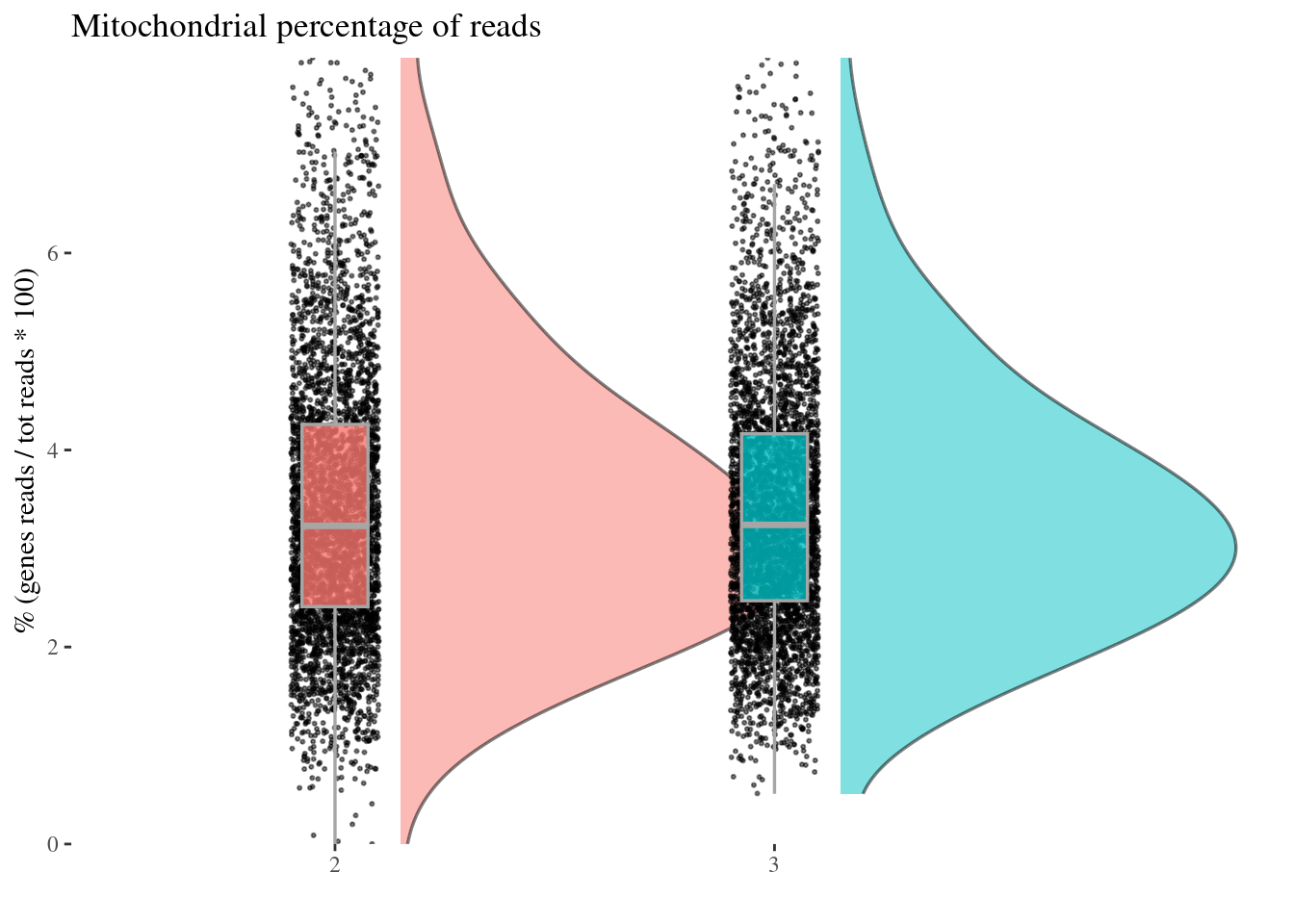
Check for spurious clusters
aRunObj <- clean(aRunObj)
c(pcaCells, pcaCellsData, genes, UDE, nu, zoomedNu) %<-%
cleanPlots(aRunObj, includePCA = TRUE)Plot PCA and Nu
plot(pcaCells)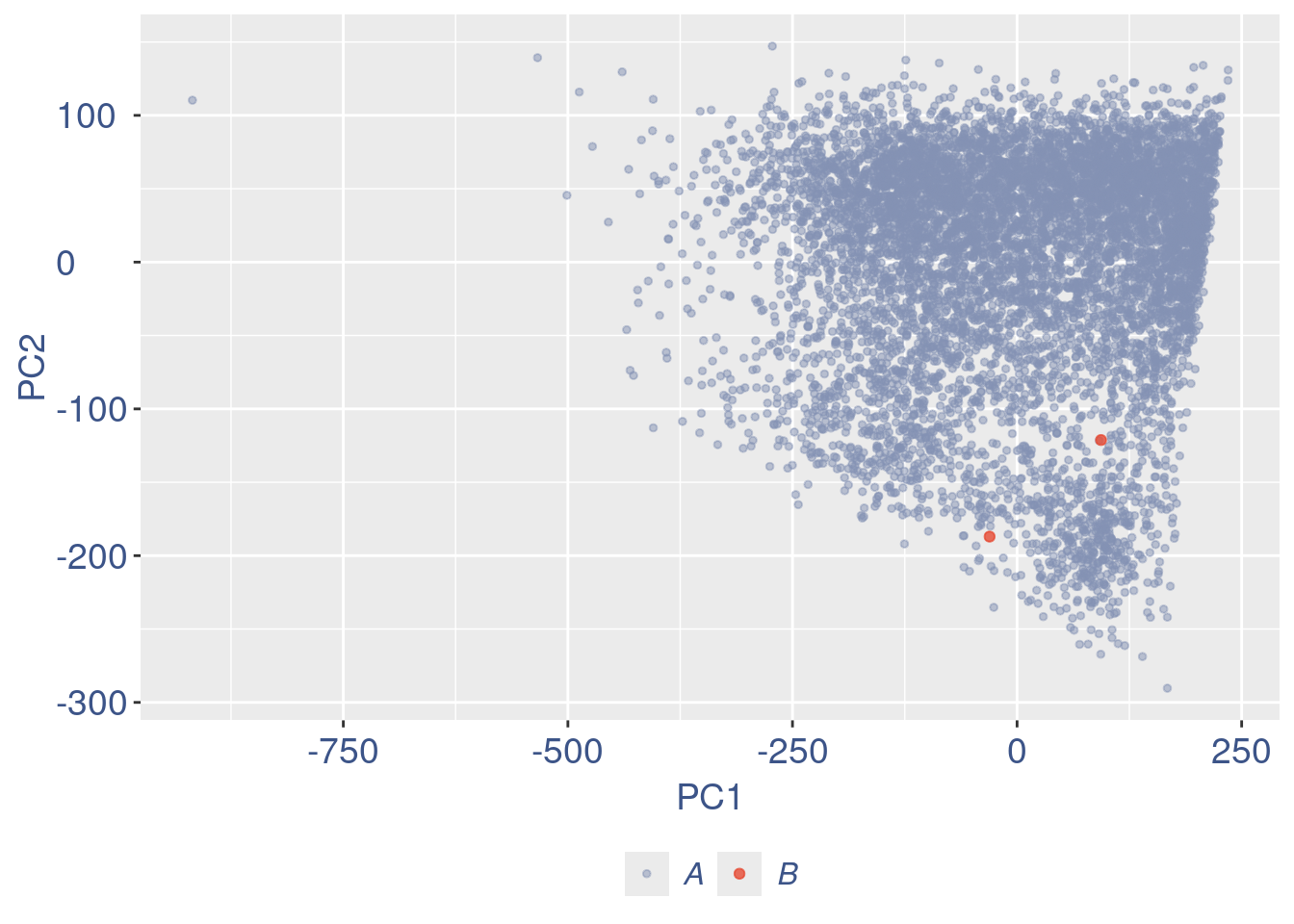
plot(UDE)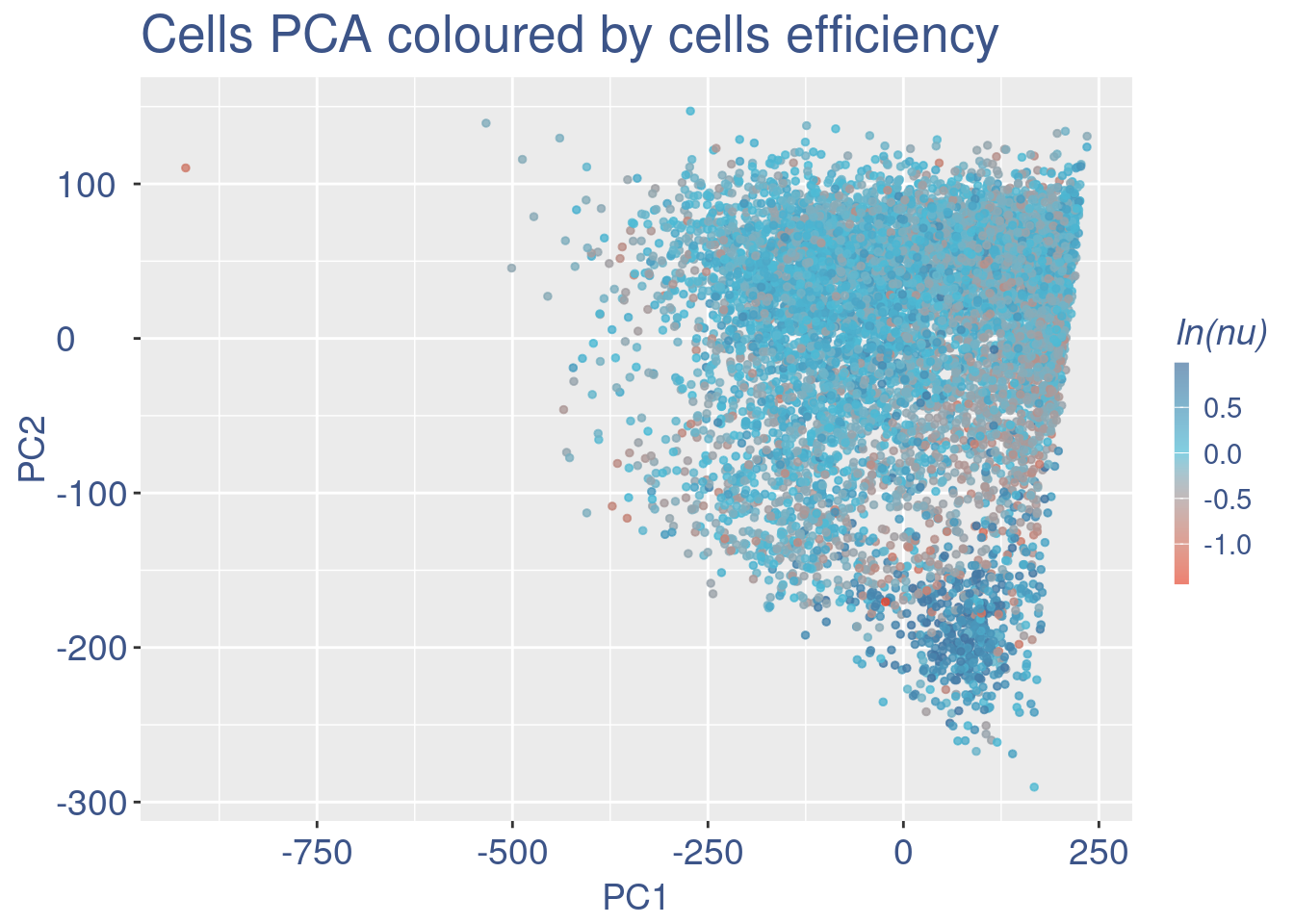
plot(pcaCellsData)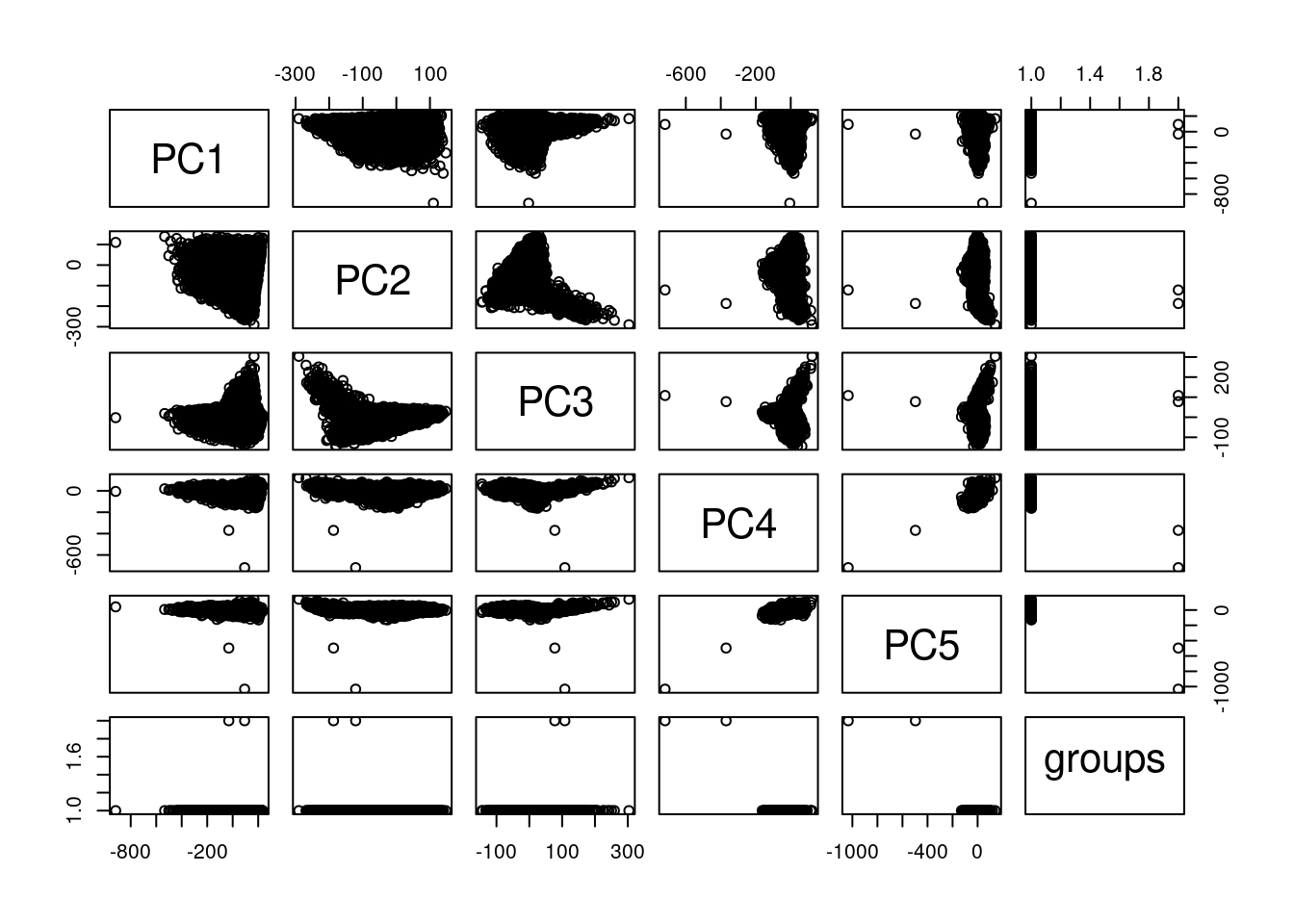
plot(genes)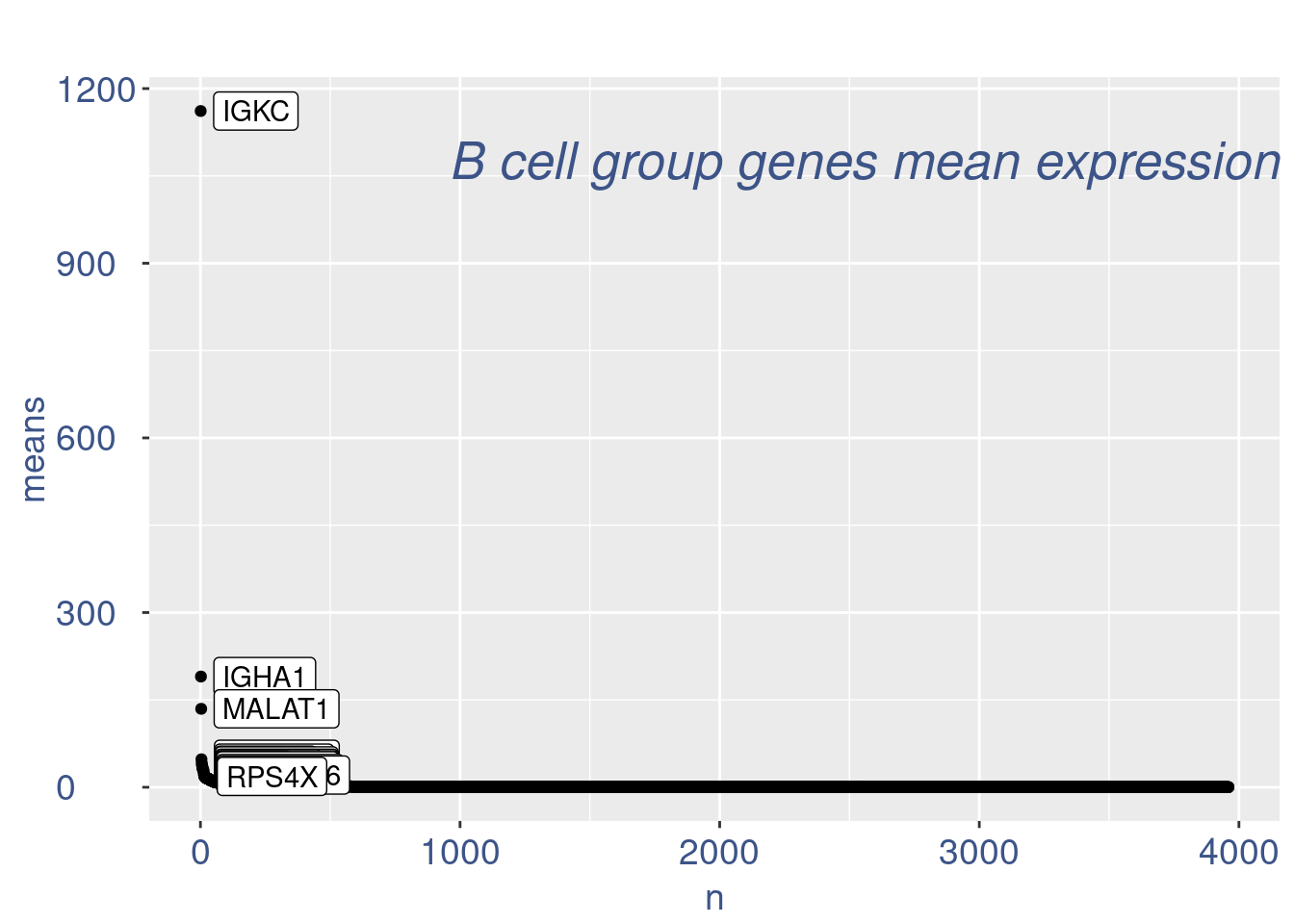
plot(nu)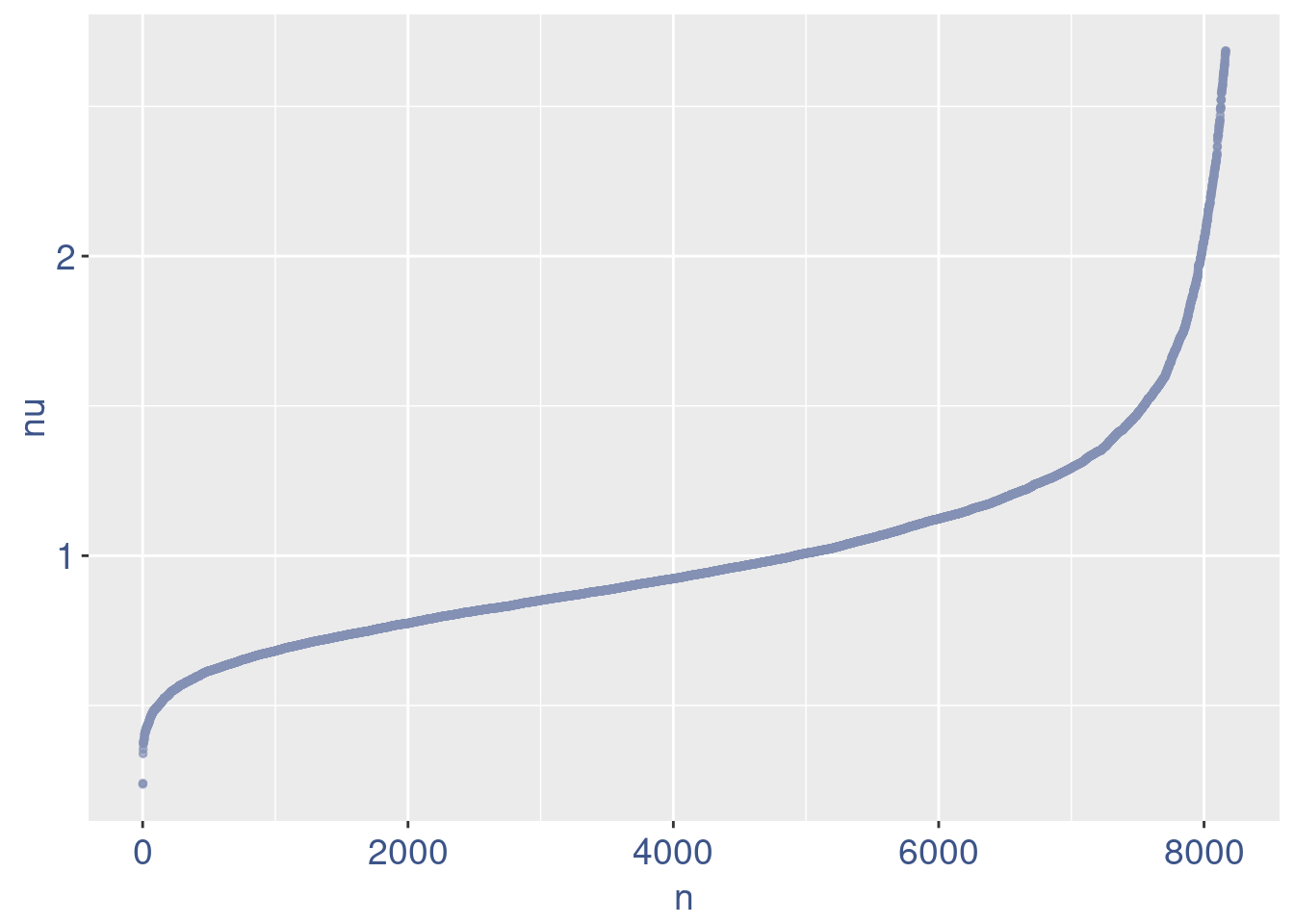
plot(zoomedNu)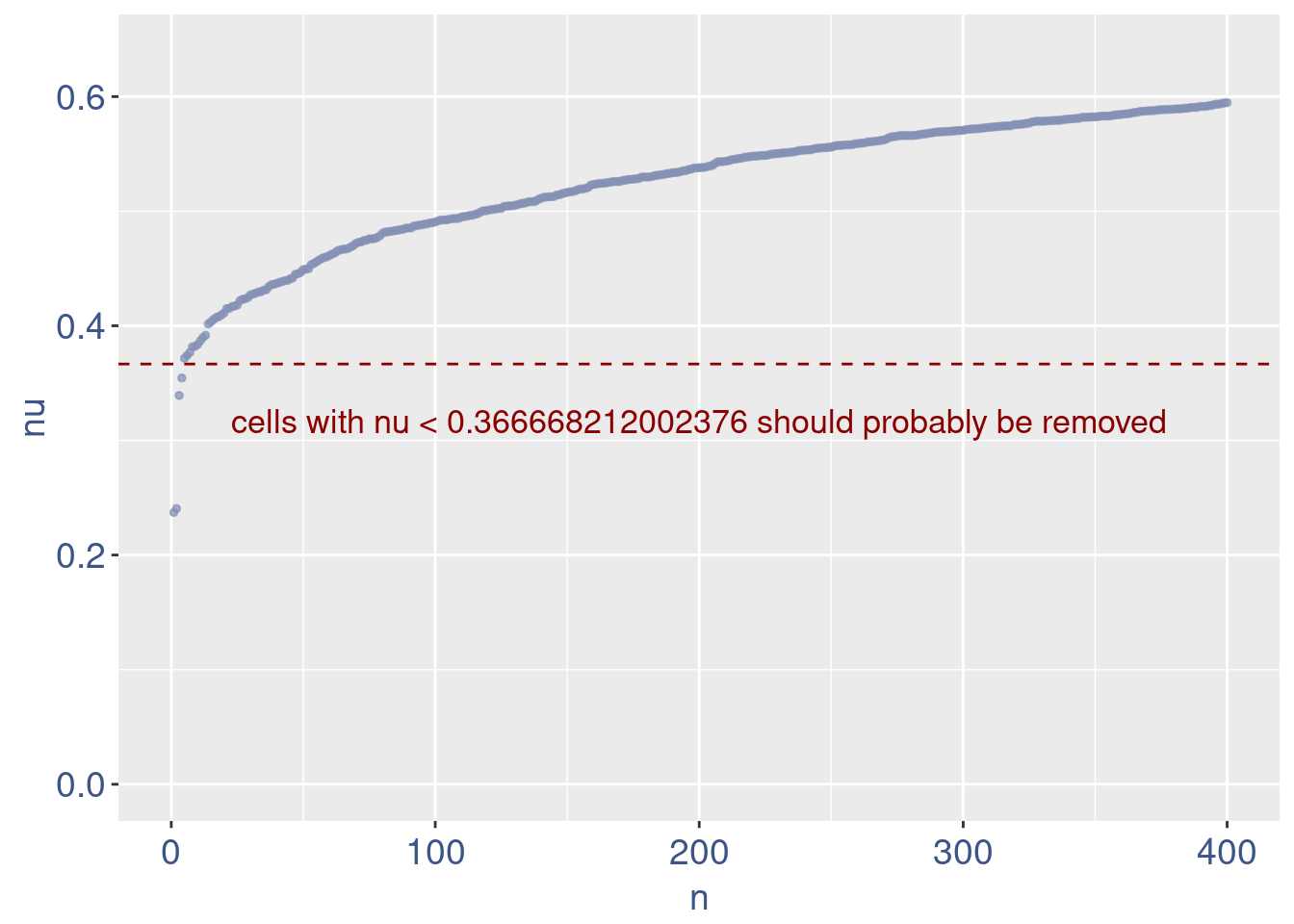
Remove cells below nu elbow point
nu <- getNu(aRunObj)
sum(nu < 0.15)[1] 0sum(nu < 0.18)[1] 0sum(nu < 0.20)[1] 0sum(nu < 0.37)[1] 4# drop above 11%
lowNuThr <- 0.37
aRunObj <-
addElementToMetaDataset(
aRunObj,
"Low Nu threshold", lowNuThr)
cells_to_rem <- rownames(pcaCellsData)[pcaCellsData[["groups"]] == "B"]
cellsToDrop <- getCells(aRunObj)[which(nu < lowNuThr)]
aRunObj <- dropGenesCells(aRunObj, cells = c(cellsToDrop,cells_to_rem))Redo all the plots
aRunObj <- clean(aRunObj)
c(pcaCells, pcaCellsData, genes, UDE, nu, zoomedNu) %<-%
cleanPlots(aRunObj, includePCA = TRUE)plot(pcaCells)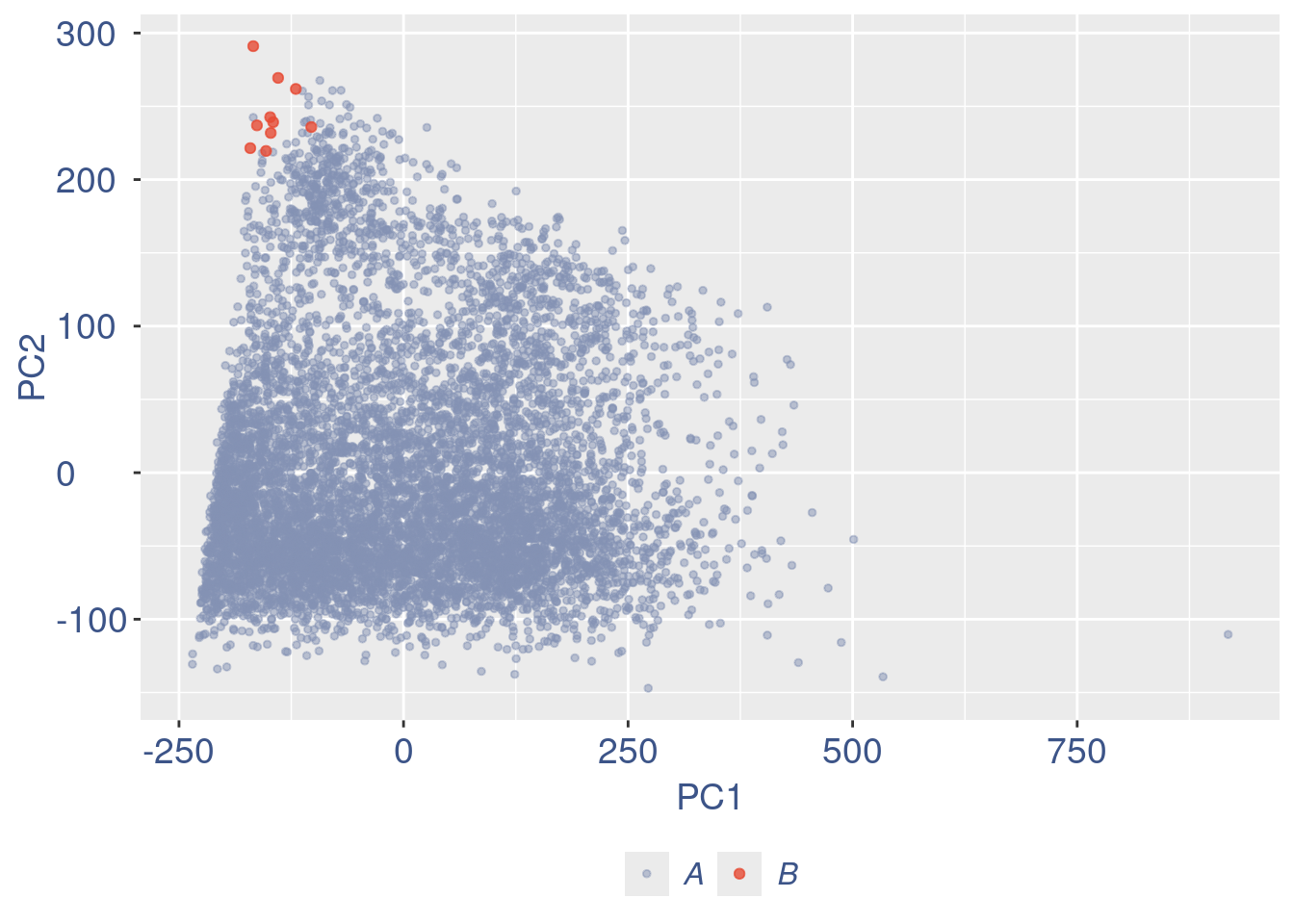
plot(UDE)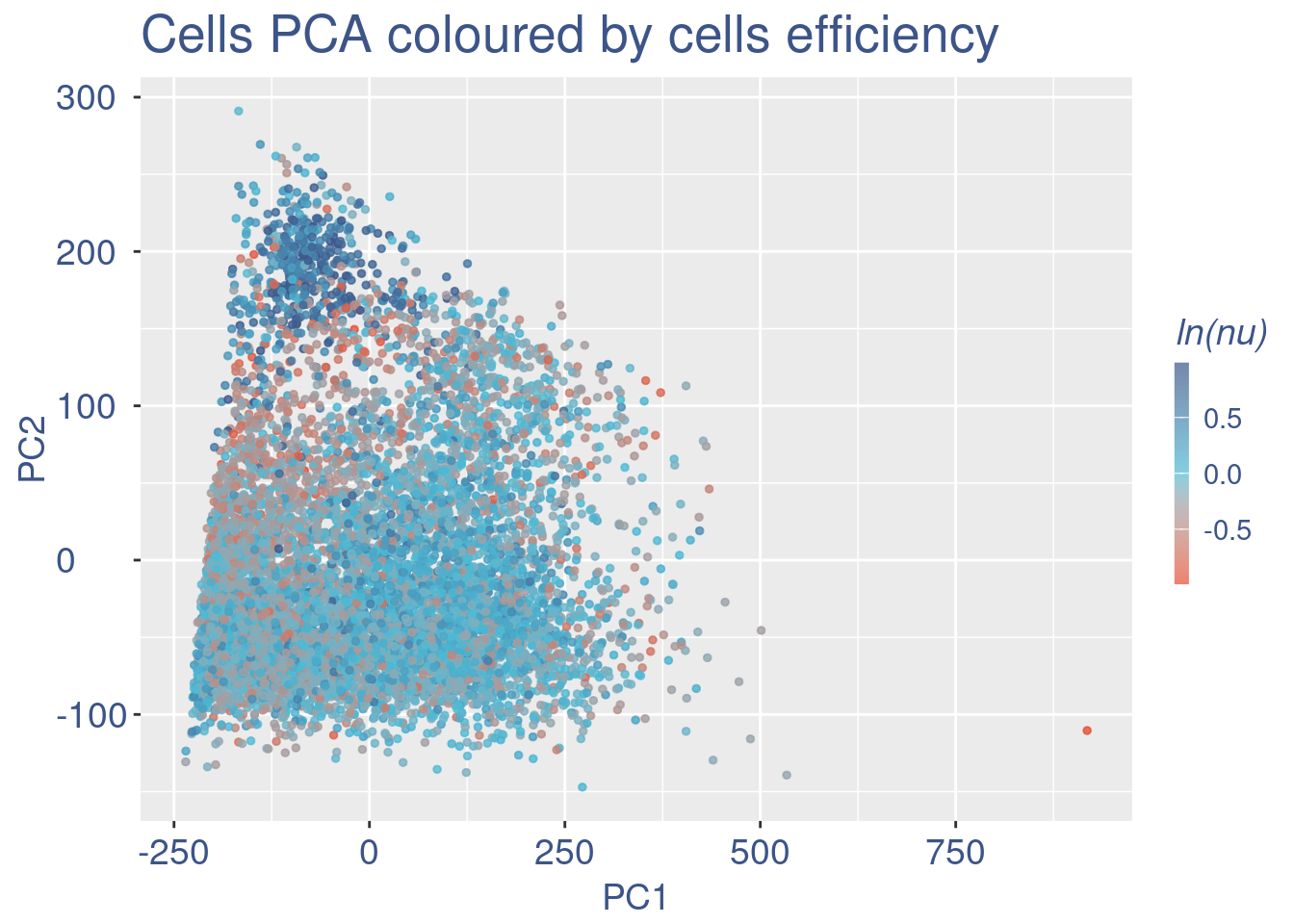
plot(pcaCellsData)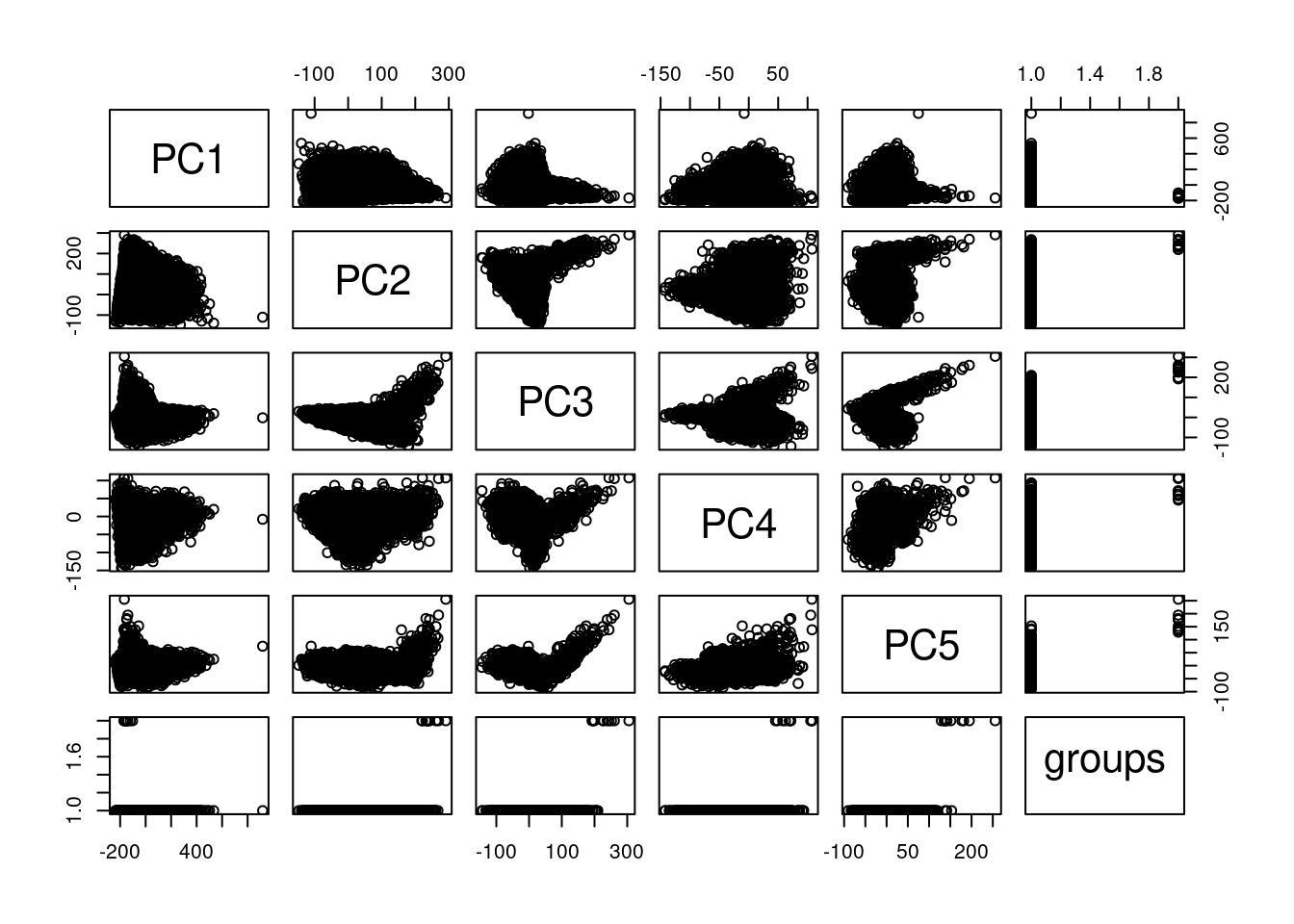
plot(genes)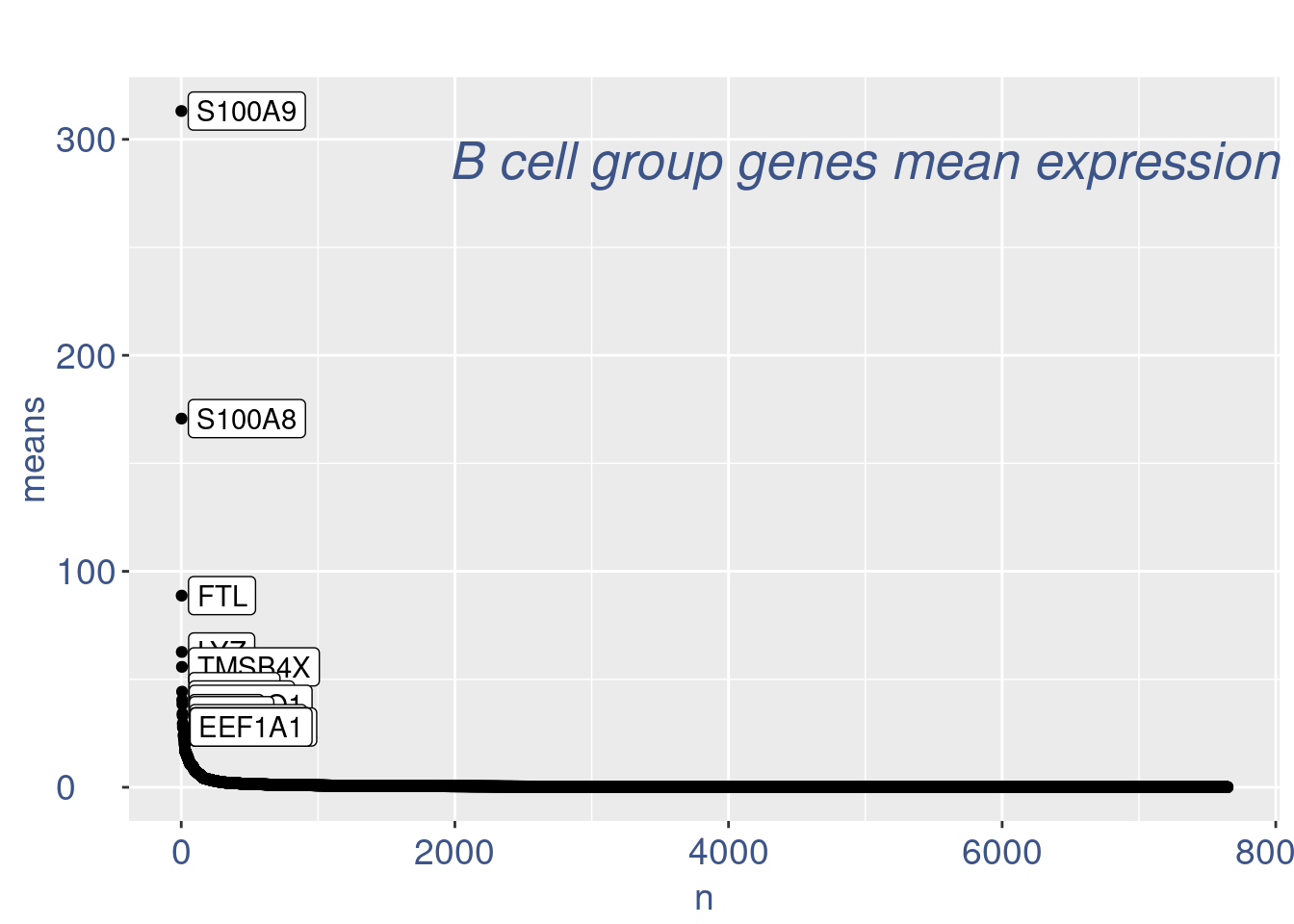
plot(nu)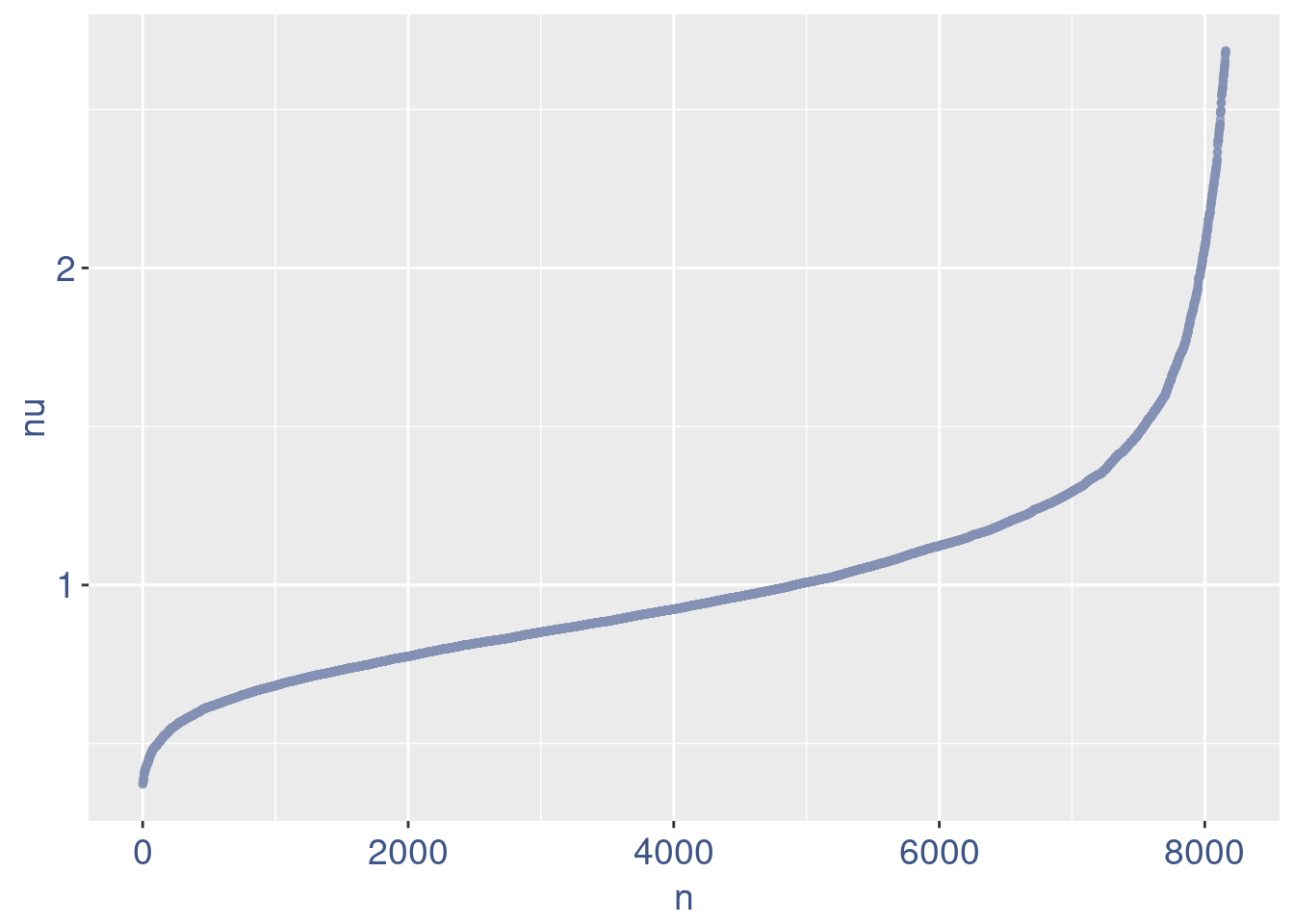
plot(zoomedNu)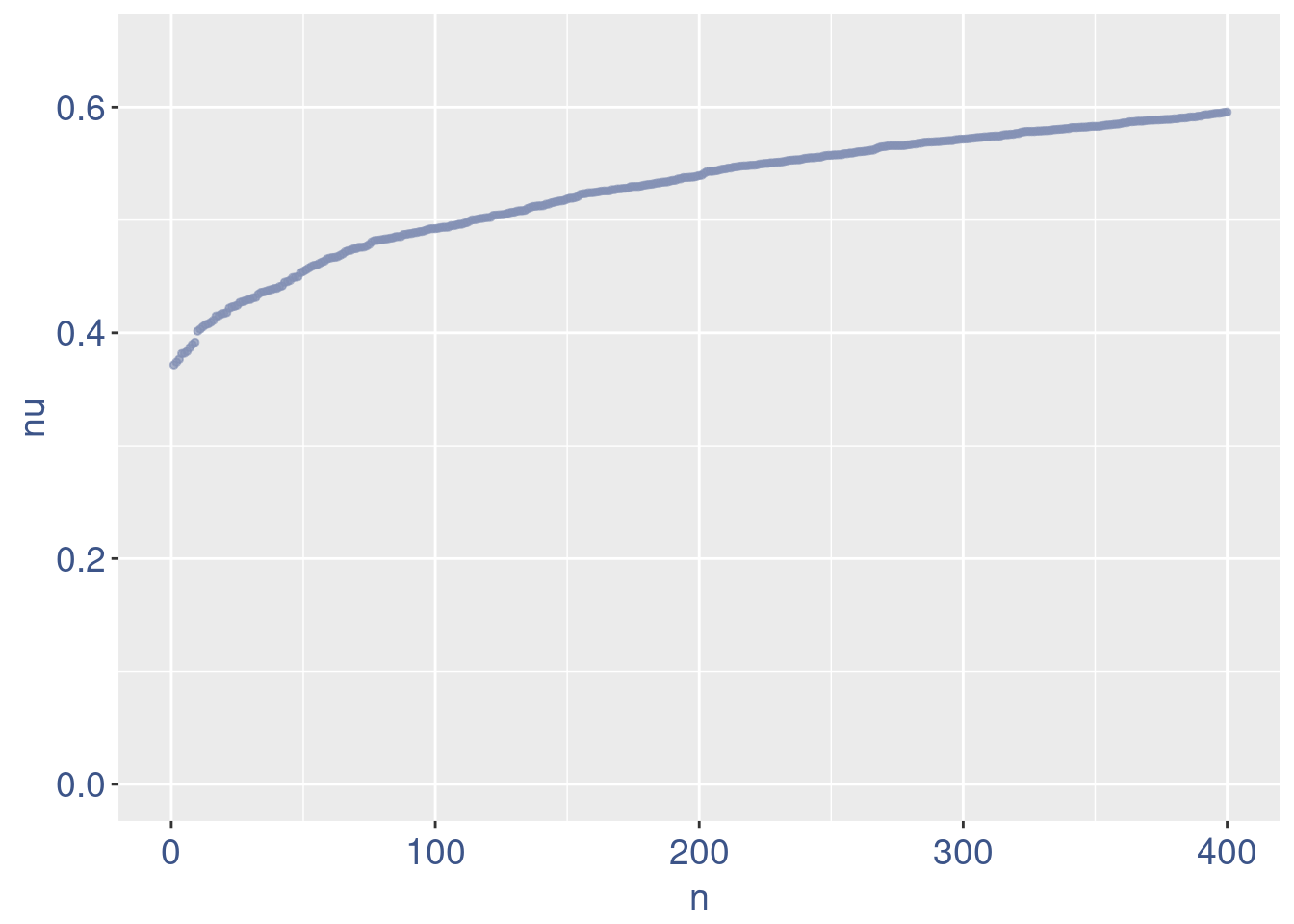
cellSizePlot(aRunObj, condName = "Participant.IDs")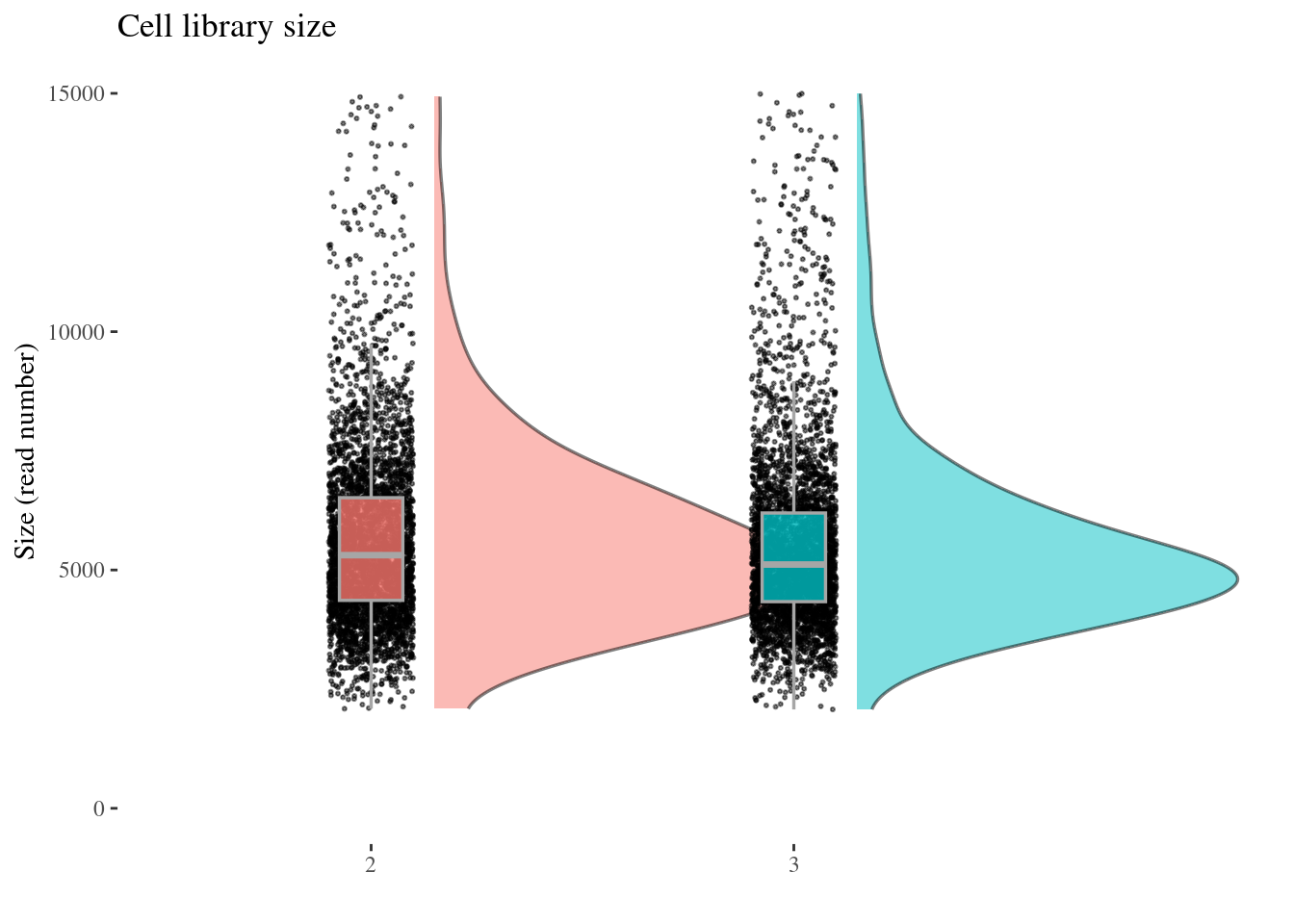
genesSizePlot(aRunObj, condName = "Participant.IDs")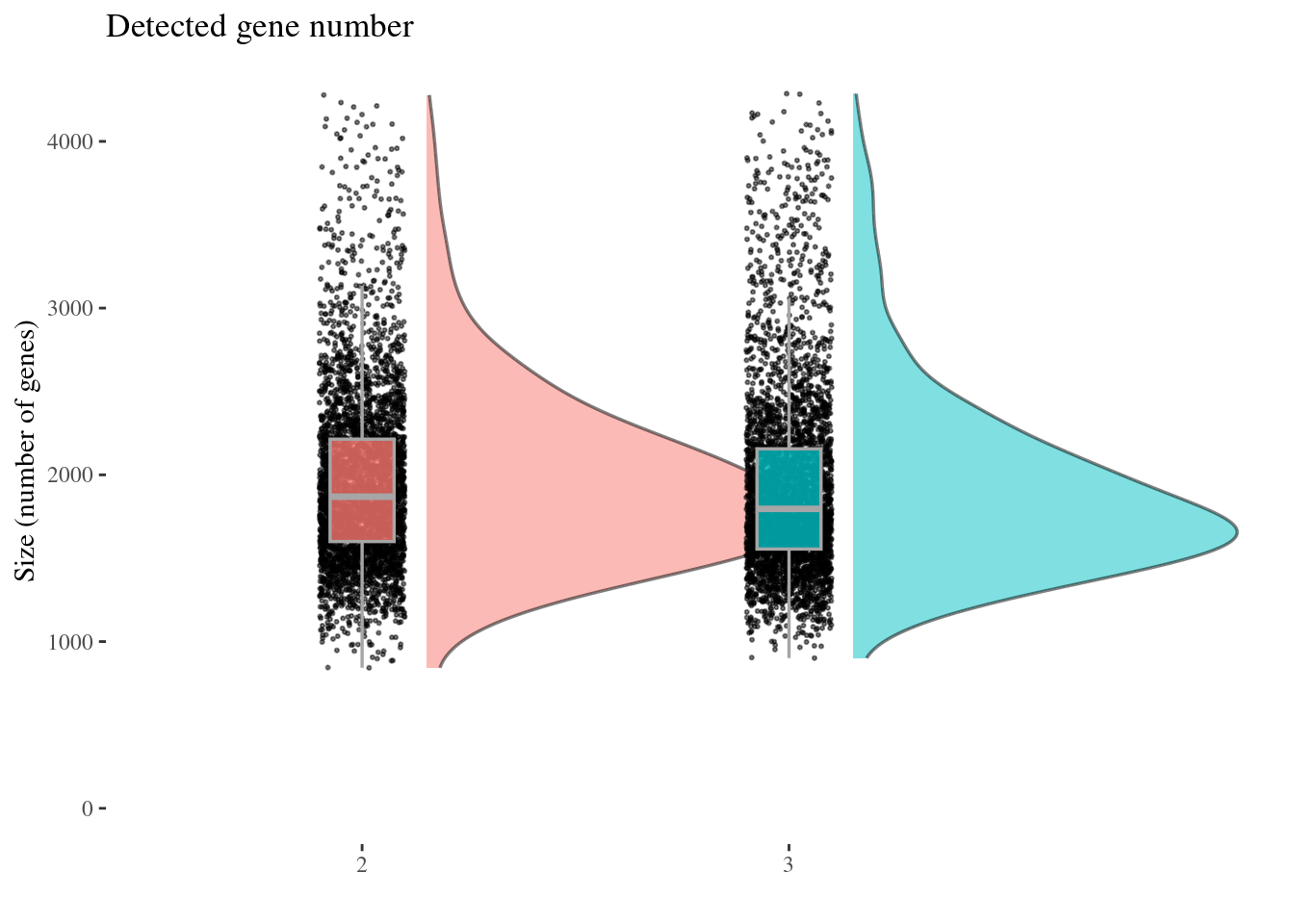
scatterPlot(aRunObj, condName = "Participant.IDs")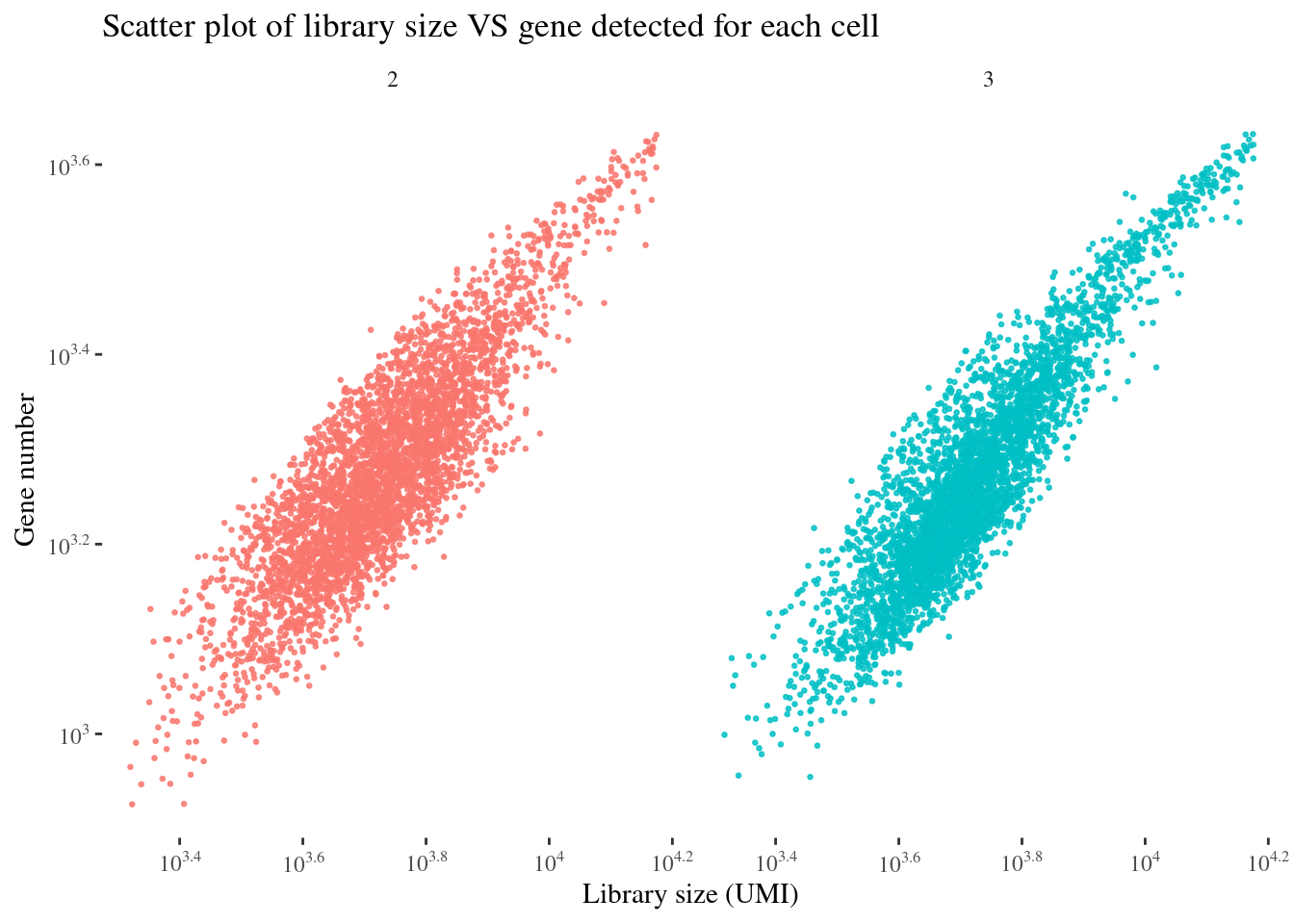
c(mitPerPlot, mitPerDf) %<-%
mitochondrialPercentagePlot(aRunObj, condName = "Participant.IDs",
genePrefix = "^MT-")
mitPerPlot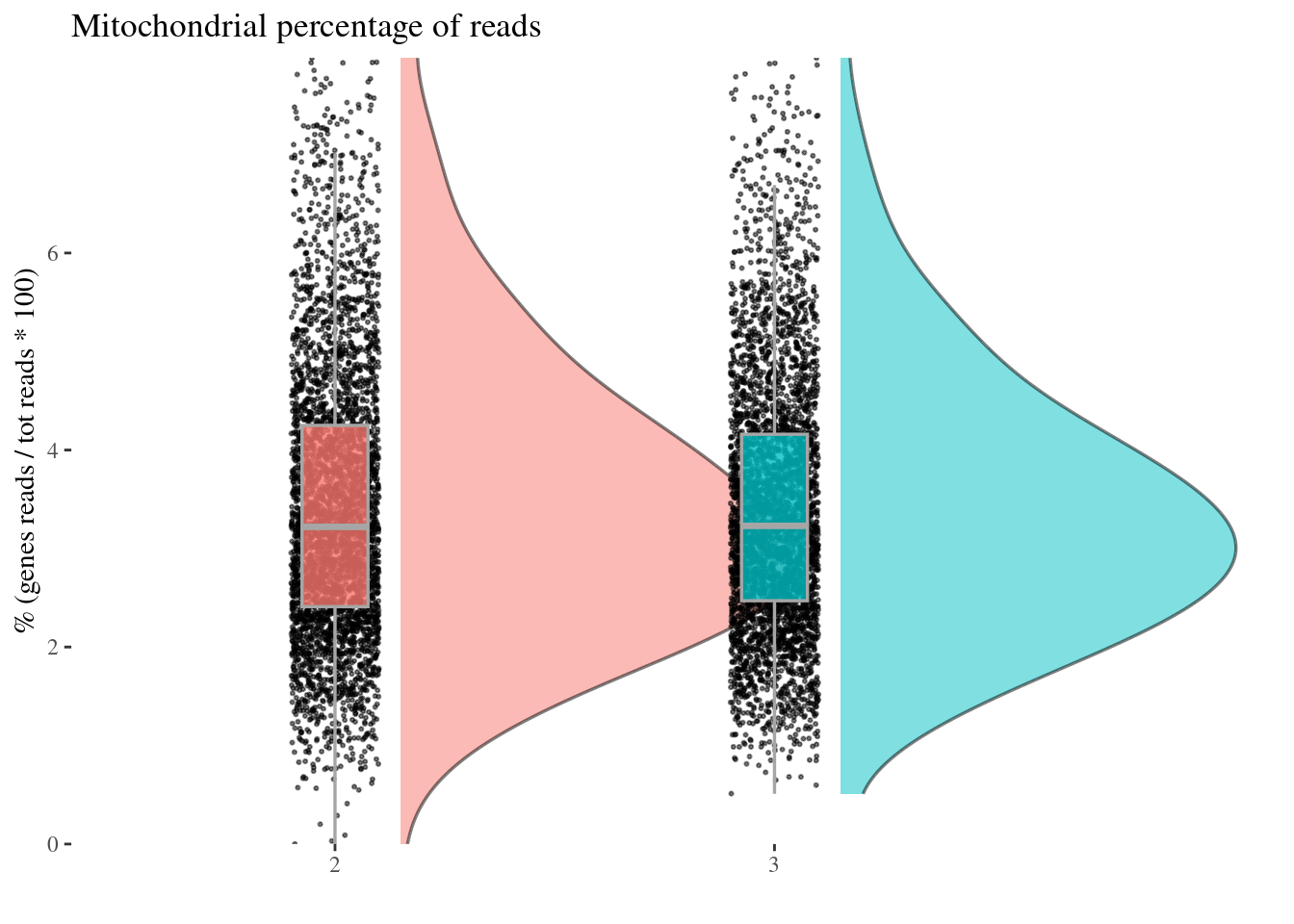
Finalize object and save
aRunObj <- proceedToCoex(aRunObj, calcCoex = FALSE, cores = 1L, saveObj = FALSE)fileNameOut <- paste0(globalCondition, "-Run_", thisRun, "-Cleaned", ".RDS")
saveRDS(aRunObj, file = file.path(outDir, fileNameOut))c2Clusters <- getClusters(aRunObj,"c2")
table(c2Clusters)c2Clusters
NK Cells B Cells Monocytes T Cells Dendritic Cells
526 431 460 6711 30 cellSizePlot(aRunObj, conditions = c2Clusters)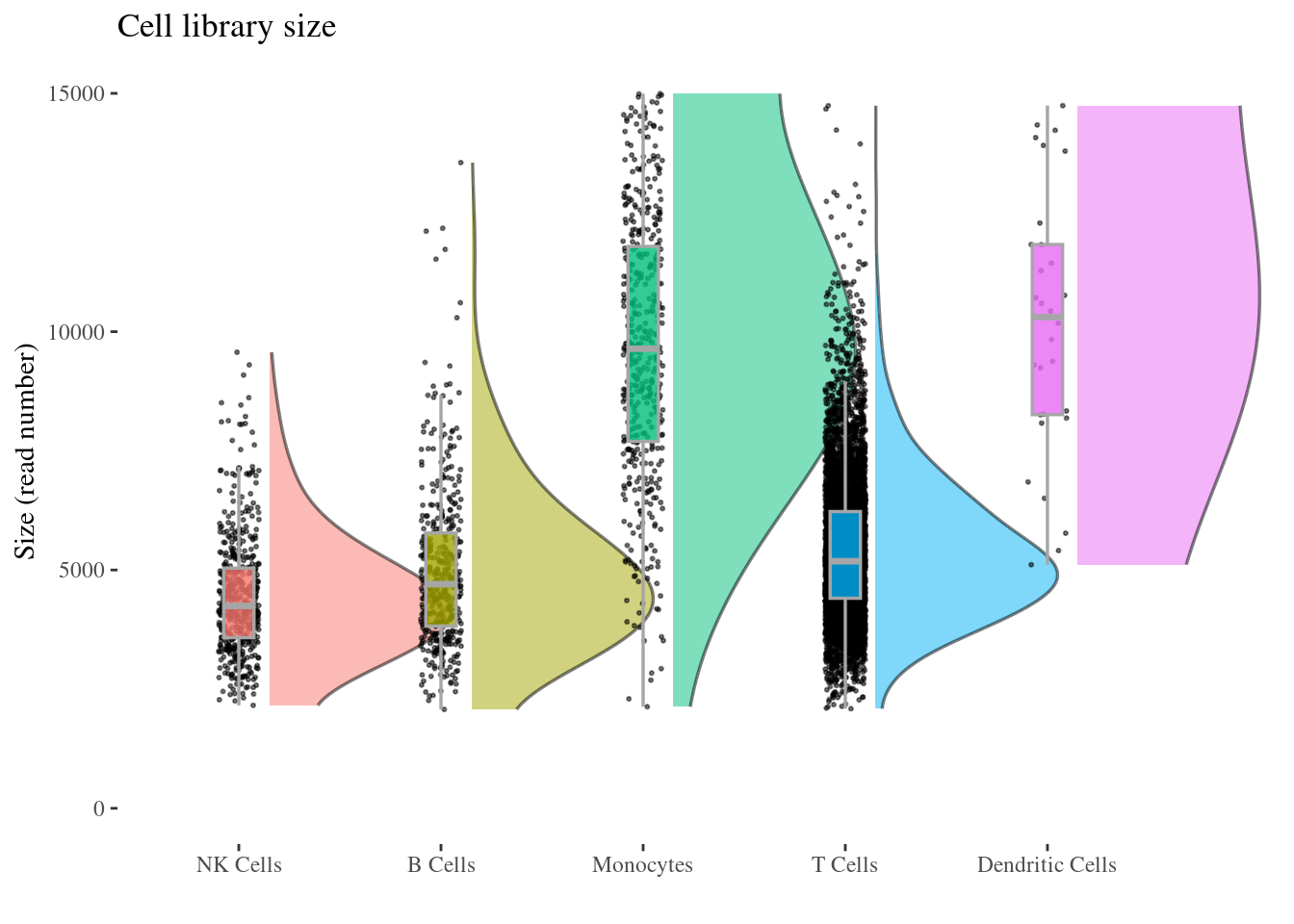
genesSizePlot(aRunObj, conditions = c2Clusters)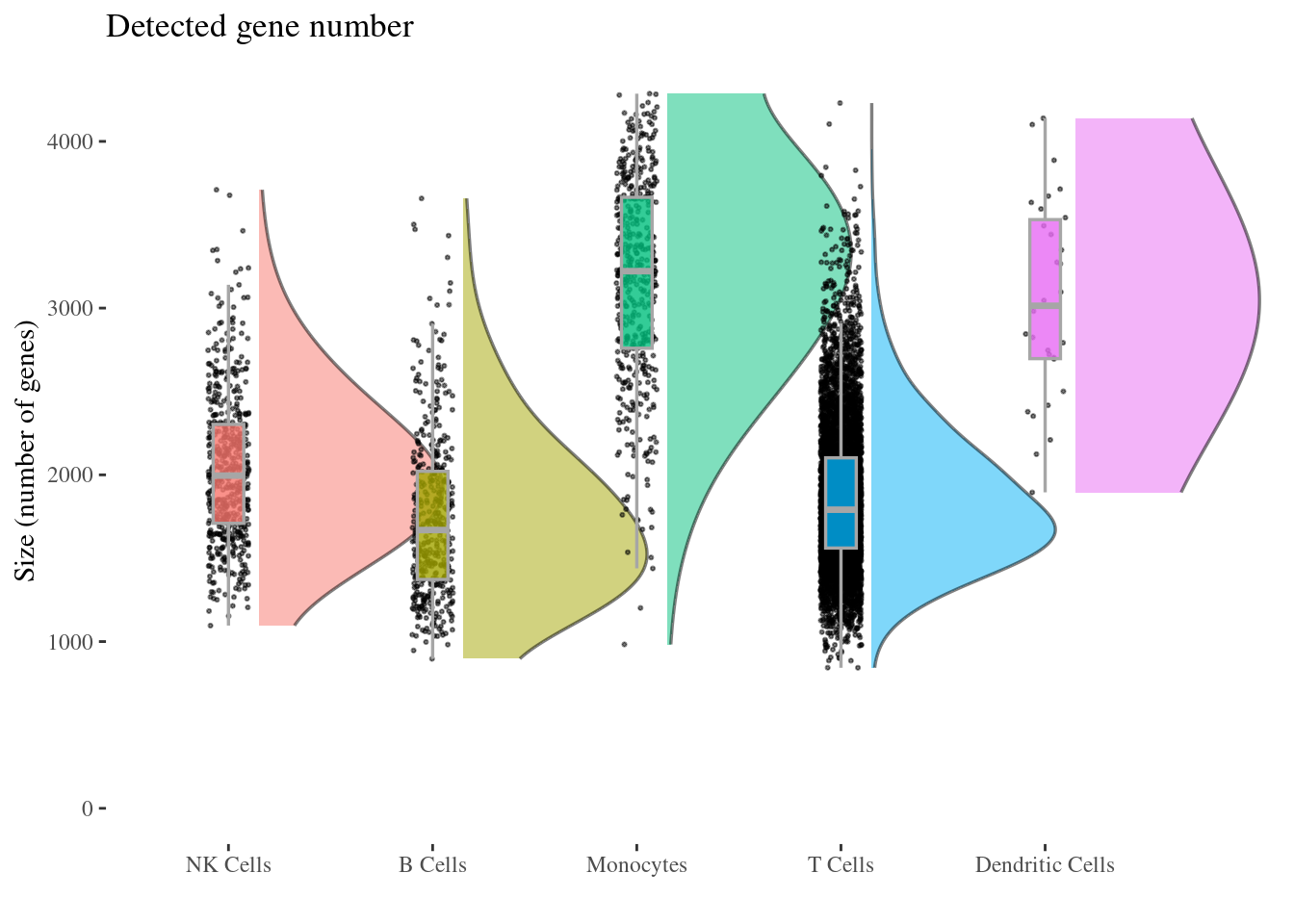
scatterPlot(aRunObj, conditions = c2Clusters)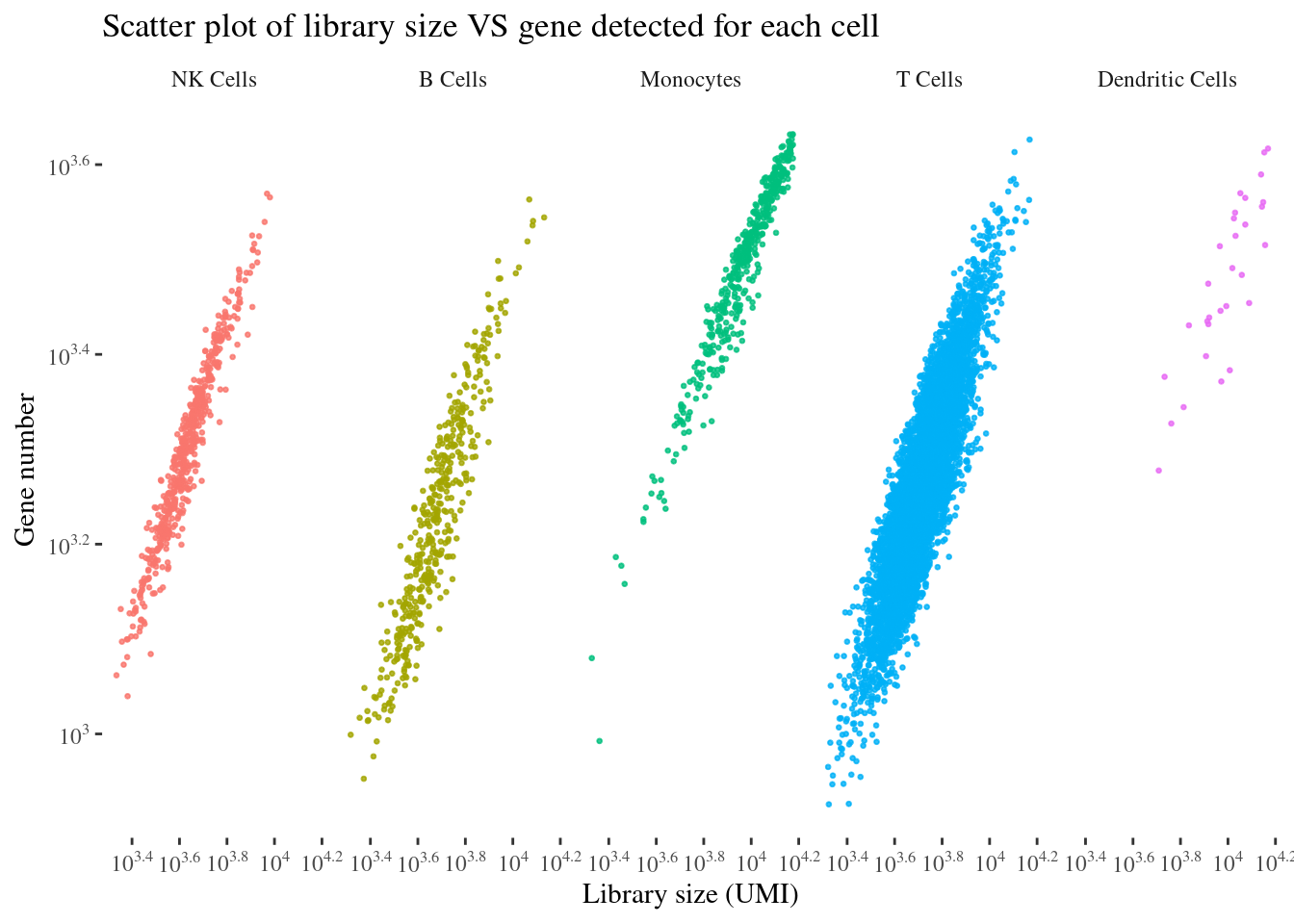
mitochondrialPercentagePlot(aRunObj, conditions = c2Clusters,
genePrefix = "^MT-")[["plot"]]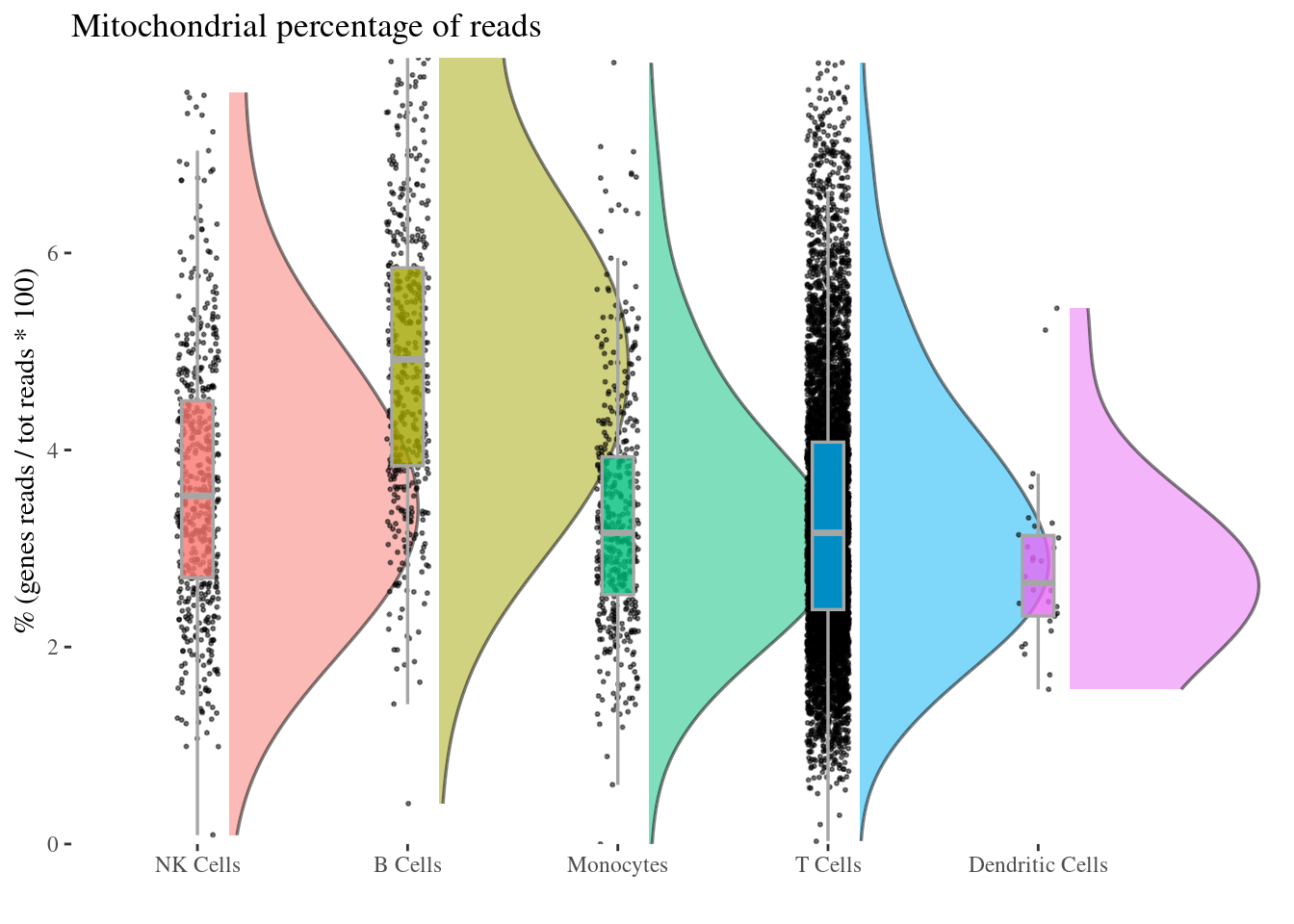
c3Clusters <- getClusters(aRunObj,"c3")
table(c3Clusters)c3Clusters
pDC CD4 T Cells
18 4607
Gamma-Delta T Cells cDC2
266 7
Intermediate Monocytes cDC3
49 4
CD56 Bright NK Cells Classical Memory B Cells
66 48
Mucosal-Associated Invariant T Cells CD56 Dim NK Cells
276 436
Classical Monocytes CD8 T Cells
314 1562
Nonclassical Monocytes Naive B Cells
97 315
asDC IgM Memory B Cells
1 62
Age-associated B Cells Adaptive NK Cells
6 24 cellSizePlot(aRunObj, conditions = c3Clusters)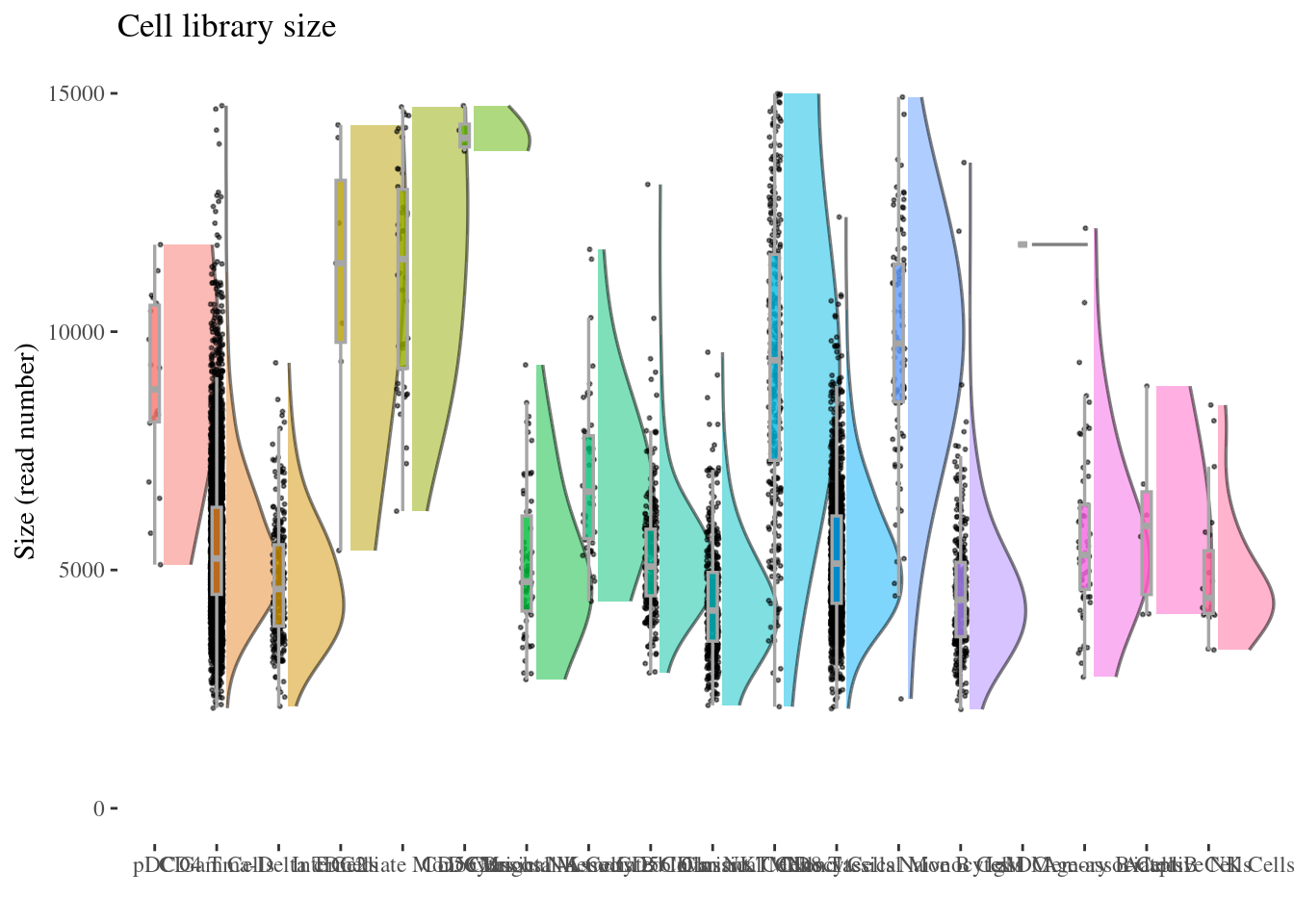
genesSizePlot(aRunObj, conditions = c3Clusters)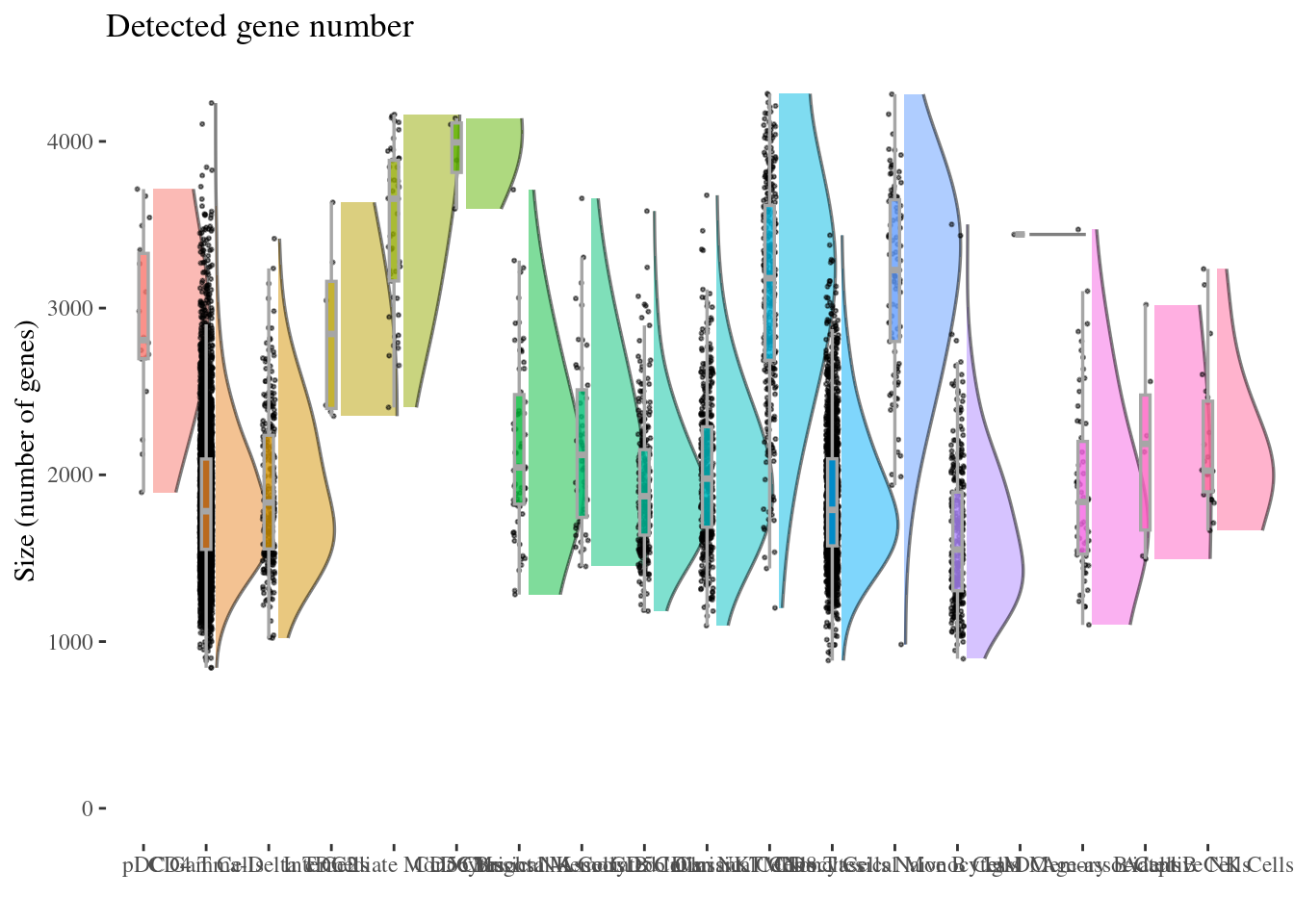
scatterPlot(aRunObj, conditions = c3Clusters)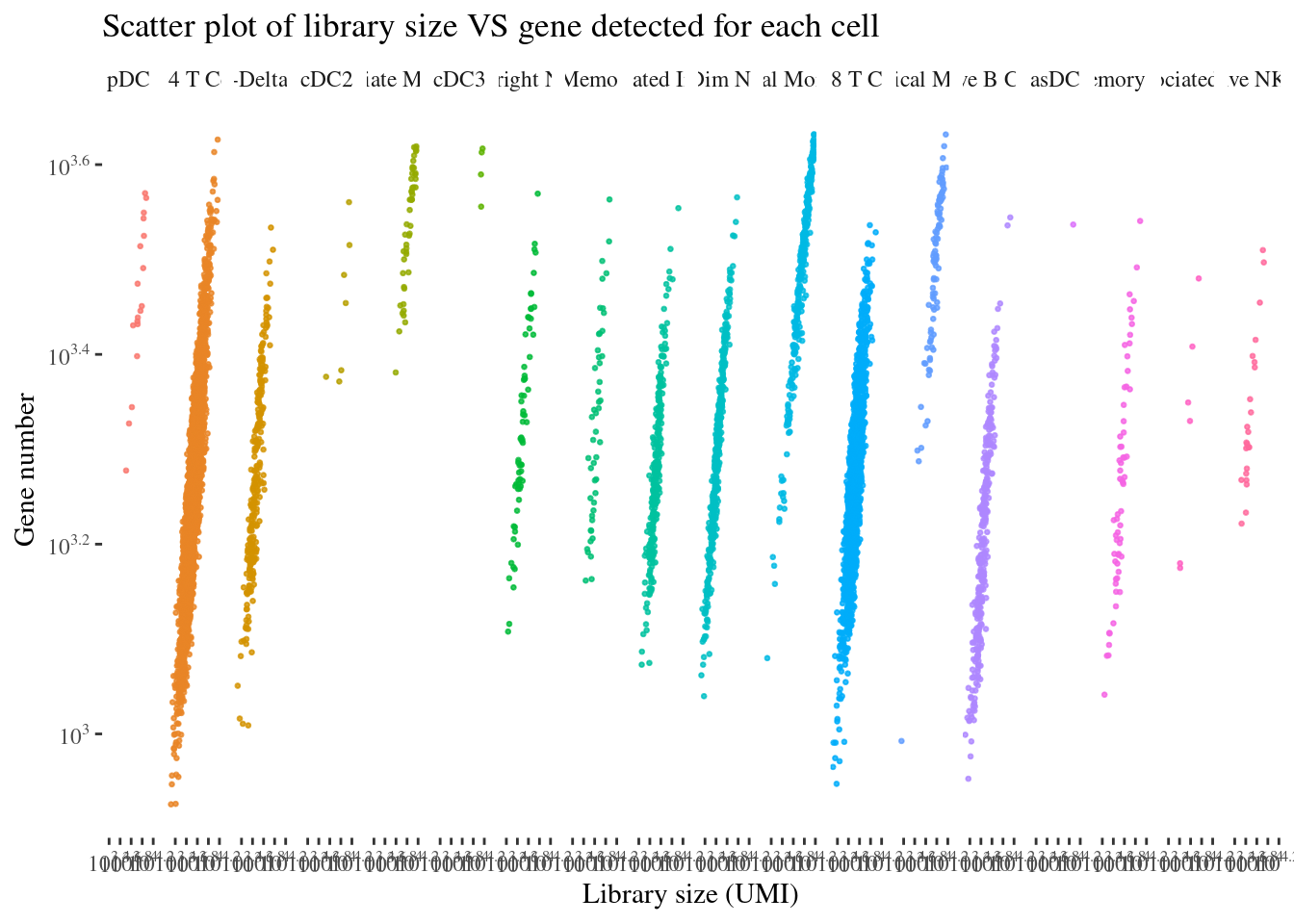
mitochondrialPercentagePlot(aRunObj, conditions = c3Clusters,
genePrefix = "^MT-")[["plot"]]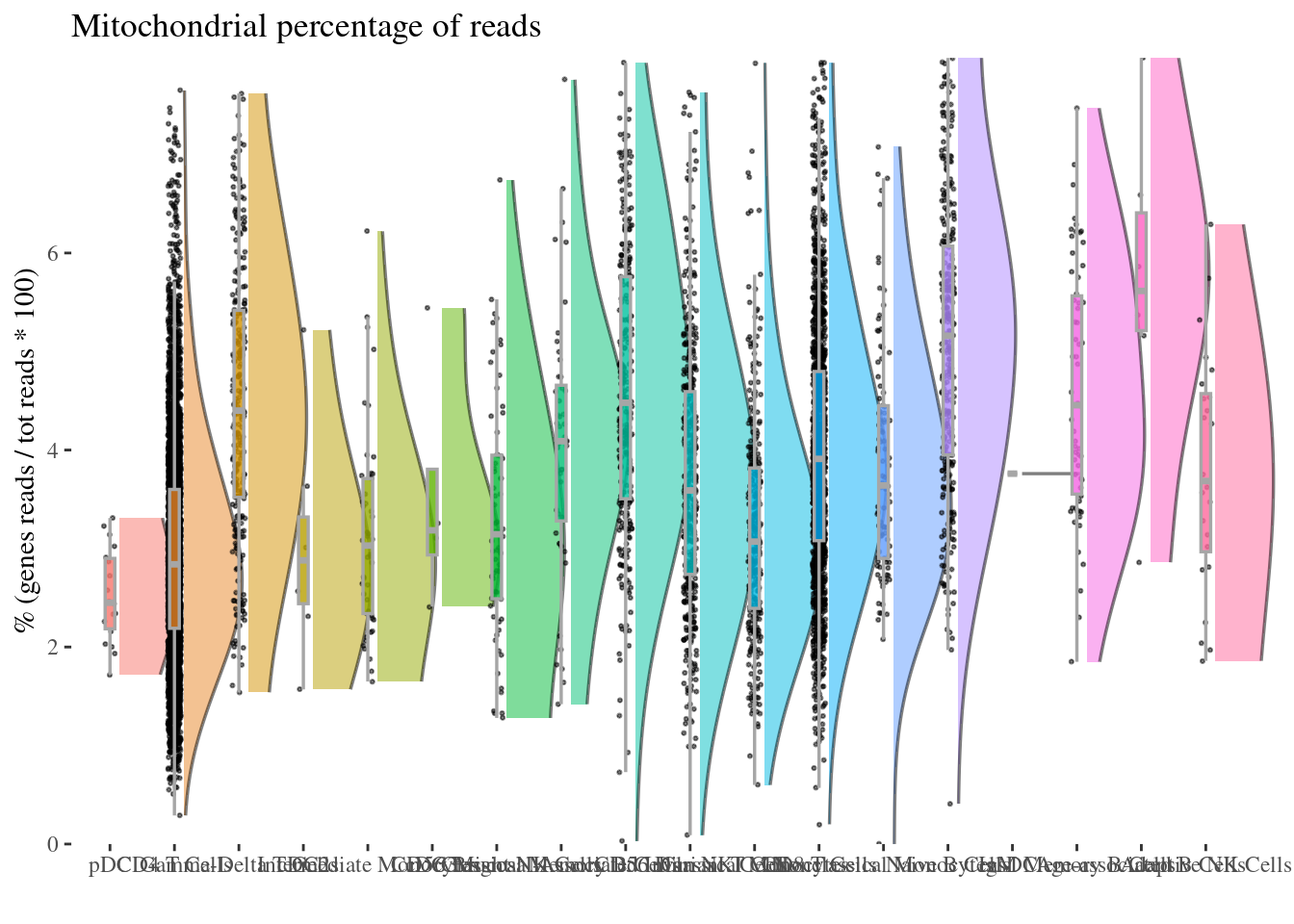
Check uniform clusters
Define check function
getClData <- function(objCOTAN, clList, cl) {
checker <- new("SimpleGDIUniformityCheck")
clName <- names(clList)[[cl]]
cells <- clList[[cl]]
res <- checkClusterUniformity(
objCOTAN, clusterName = clName, cells = cells,
checker = checker, cores = 3L, saveObj = FALSE
)
return(res)
}Create new clusterization from C3 and SampleID
newCl <- paste(niceFactorLevels(getCondition(aRunObj, condName = "Sample.IDs")),
getClusters(aRunObj, clName = "c3"), sep = "_")
names(newCl) <- getCells(aRunObj)
newCl <- base::as.factor(newCl)
aRunObj <- addClusterization(aRunObj, clName = "Sample_C3", clusters = newCl)
rm(newCl)
narrowC3Clusters <- getClusters(aRunObj, clName = "Sample_C3")
sort(table(narrowC3Clusters), decreasing = TRUE)narrowC3Clusters
095_CD4 T Cells
2478
424_CD4 T Cells
2129
424_CD8 T Cells
845
095_CD8 T Cells
717
424_CD56 Dim NK Cells
267
095_Classical Monocytes
228
424_Mucosal-Associated Invariant T Cells
176
095_CD56 Dim NK Cells
169
424_Naive B Cells
159
095_Naive B Cells
156
424_Gamma-Delta T Cells
154
095_Gamma-Delta T Cells
112
095_Mucosal-Associated Invariant T Cells
100
424_Classical Monocytes
86
095_Nonclassical Monocytes
69
424_IgM Memory B Cells
42
424_Classical Memory B Cells
39
095_CD56 Bright NK Cells
34
424_CD56 Bright NK Cells
32
424_Nonclassical Monocytes
28
095_Intermediate Monocytes
27
424_Intermediate Monocytes
22
095_IgM Memory B Cells
20
424_Adaptive NK Cells
18
424_pDC
10
095_Classical Memory B Cells
9
095_pDC
8
095_Adaptive NK Cells
6
424_Age-associated B Cells
5
095_cDC2
4
424_cDC2
3
095_cDC3
2
424_cDC3
2
095_Age-associated B Cells
1
095_asDC
1 Pick largest clusters
largeClNames <- names(which(table(narrowC3Clusters) >= 50))
largeCl <- narrowC3Clusters[narrowC3Clusters %in% largeClNames]
largeCl <- toClustersList(largeCl)GDI checks for larger clusters
lengths(largeCl) 095_CD4 T Cells
2478
095_CD56 Dim NK Cells
169
095_CD8 T Cells
717
095_Classical Monocytes
228
095_Gamma-Delta T Cells
112
095_Mucosal-Associated Invariant T Cells
100
095_Naive B Cells
156
095_Nonclassical Monocytes
69
424_CD4 T Cells
2129
424_CD56 Dim NK Cells
267
424_CD8 T Cells
845
424_Classical Monocytes
86
424_Gamma-Delta T Cells
154
424_Mucosal-Associated Invariant T Cells
176
424_Naive B Cells
159 resNarrC3Data <- lapply(seq_along(largeCl), FUN = getClData,
objCOTAN = aRunObj, clList = largeCl)
names(resNarrC3Data) <- names(largeCl)
narrowC3ClData <- checkersToDF(resNarrC3Data)
rownames(narrowC3ClData) <- names(largeCl)
narrowC3ClData class
095_CD4 T Cells SimpleGDIUniformityCheck
095_CD56 Dim NK Cells SimpleGDIUniformityCheck
095_CD8 T Cells SimpleGDIUniformityCheck
095_Classical Monocytes SimpleGDIUniformityCheck
095_Gamma-Delta T Cells SimpleGDIUniformityCheck
095_Mucosal-Associated Invariant T Cells SimpleGDIUniformityCheck
095_Naive B Cells SimpleGDIUniformityCheck
095_Nonclassical Monocytes SimpleGDIUniformityCheck
424_CD4 T Cells SimpleGDIUniformityCheck
424_CD56 Dim NK Cells SimpleGDIUniformityCheck
424_CD8 T Cells SimpleGDIUniformityCheck
424_Classical Monocytes SimpleGDIUniformityCheck
424_Gamma-Delta T Cells SimpleGDIUniformityCheck
424_Mucosal-Associated Invariant T Cells SimpleGDIUniformityCheck
424_Naive B Cells SimpleGDIUniformityCheck
check.isCheckAbove check.GDIThreshold
095_CD4 T Cells FALSE 1.4
095_CD56 Dim NK Cells FALSE 1.4
095_CD8 T Cells FALSE 1.4
095_Classical Monocytes FALSE 1.4
095_Gamma-Delta T Cells FALSE 1.4
095_Mucosal-Associated Invariant T Cells FALSE 1.4
095_Naive B Cells FALSE 1.4
095_Nonclassical Monocytes FALSE 1.4
424_CD4 T Cells FALSE 1.4
424_CD56 Dim NK Cells FALSE 1.4
424_CD8 T Cells FALSE 1.4
424_Classical Monocytes FALSE 1.4
424_Gamma-Delta T Cells FALSE 1.4
424_Mucosal-Associated Invariant T Cells FALSE 1.4
424_Naive B Cells FALSE 1.4
check.maxRatioBeyond
095_CD4 T Cells 0.01
095_CD56 Dim NK Cells 0.01
095_CD8 T Cells 0.01
095_Classical Monocytes 0.01
095_Gamma-Delta T Cells 0.01
095_Mucosal-Associated Invariant T Cells 0.01
095_Naive B Cells 0.01
095_Nonclassical Monocytes 0.01
424_CD4 T Cells 0.01
424_CD56 Dim NK Cells 0.01
424_CD8 T Cells 0.01
424_Classical Monocytes 0.01
424_Gamma-Delta T Cells 0.01
424_Mucosal-Associated Invariant T Cells 0.01
424_Naive B Cells 0.01
check.maxRankBeyond
095_CD4 T Cells 0
095_CD56 Dim NK Cells 0
095_CD8 T Cells 0
095_Classical Monocytes 0
095_Gamma-Delta T Cells 0
095_Mucosal-Associated Invariant T Cells 0
095_Naive B Cells 0
095_Nonclassical Monocytes 0
424_CD4 T Cells 0
424_CD56 Dim NK Cells 0
424_CD8 T Cells 0
424_Classical Monocytes 0
424_Gamma-Delta T Cells 0
424_Mucosal-Associated Invariant T Cells 0
424_Naive B Cells 0
check.fractionBeyond
095_CD4 T Cells 0.410118632
095_CD56 Dim NK Cells 0.036571045
095_CD8 T Cells 0.181047489
095_Classical Monocytes 0.027420871
095_Gamma-Delta T Cells 0.022672754
095_Mucosal-Associated Invariant T Cells 0.001211338
095_Naive B Cells 0.081778266
095_Nonclassical Monocytes 0.010109997
424_CD4 T Cells 0.435352338
424_CD56 Dim NK Cells 0.084106959
424_CD8 T Cells 0.307495989
424_Classical Monocytes 0.002347969
424_Gamma-Delta T Cells 0.055401454
424_Mucosal-Associated Invariant T Cells 0.066509712
424_Naive B Cells 0.055647775
check.thresholdRank
095_CD4 T Cells 0
095_CD56 Dim NK Cells 0
095_CD8 T Cells 0
095_Classical Monocytes 0
095_Gamma-Delta T Cells 0
095_Mucosal-Associated Invariant T Cells 0
095_Naive B Cells 0
095_Nonclassical Monocytes 0
424_CD4 T Cells 0
424_CD56 Dim NK Cells 0
424_CD8 T Cells 0
424_Classical Monocytes 0
424_Gamma-Delta T Cells 0
424_Mucosal-Associated Invariant T Cells 0
424_Naive B Cells 0
check.quantileAtRatio
095_CD4 T Cells 2.353247
095_CD56 Dim NK Cells 1.465200
095_CD8 T Cells 1.775139
095_Classical Monocytes 1.447324
095_Gamma-Delta T Cells 1.443751
095_Mucosal-Associated Invariant T Cells 1.326855
095_Naive B Cells 1.599132
095_Nonclassical Monocytes 1.400252
424_CD4 T Cells 2.403522
424_CD56 Dim NK Cells 1.536310
424_CD8 T Cells 2.099888
424_Classical Monocytes 1.354043
424_Gamma-Delta T Cells 1.519138
424_Mucosal-Associated Invariant T Cells 1.523826
424_Naive B Cells 1.539789
check.quantileAtRank isUniform
095_CD4 T Cells NaN FALSE
095_CD56 Dim NK Cells NaN FALSE
095_CD8 T Cells NaN FALSE
095_Classical Monocytes NaN FALSE
095_Gamma-Delta T Cells NaN FALSE
095_Mucosal-Associated Invariant T Cells NaN TRUE
095_Naive B Cells NaN FALSE
095_Nonclassical Monocytes NaN FALSE
424_CD4 T Cells NaN FALSE
424_CD56 Dim NK Cells NaN FALSE
424_CD8 T Cells NaN FALSE
424_Classical Monocytes NaN TRUE
424_Gamma-Delta T Cells NaN FALSE
424_Mucosal-Associated Invariant T Cells NaN FALSE
424_Naive B Cells NaN FALSE
clusterSize
095_CD4 T Cells 2478
095_CD56 Dim NK Cells 169
095_CD8 T Cells 717
095_Classical Monocytes 228
095_Gamma-Delta T Cells 112
095_Mucosal-Associated Invariant T Cells 100
095_Naive B Cells 156
095_Nonclassical Monocytes 69
424_CD4 T Cells 2129
424_CD56 Dim NK Cells 267
424_CD8 T Cells 845
424_Classical Monocytes 86
424_Gamma-Delta T Cells 154
424_Mucosal-Associated Invariant T Cells 176
424_Naive B Cells 159shiftToUnif <- lapply(resNarrC3Data, calculateThresholdShiftToUniformity)
shiftToUnif$`095_CD4 T Cells`
[1] 0.9532475
$`095_CD56 Dim NK Cells`
[1] 0.06519974
$`095_CD8 T Cells`
[1] 0.3751387
$`095_Classical Monocytes`
[1] 0.04732434
$`095_Gamma-Delta T Cells`
[1] 0.04375143
$`095_Mucosal-Associated Invariant T Cells`
[1] -0.07314485
$`095_Naive B Cells`
[1] 0.1991318
$`095_Nonclassical Monocytes`
[1] 0.0002517048
$`424_CD4 T Cells`
[1] 1.003522
$`424_CD56 Dim NK Cells`
[1] 0.1363103
$`424_CD8 T Cells`
[1] 0.6998877
$`424_Classical Monocytes`
[1] -0.04595663
$`424_Gamma-Delta T Cells`
[1] 0.1191376
$`424_Mucosal-Associated Invariant T Cells`
[1] 0.1238262
$`424_Naive B Cells`
[1] 0.1397889Save check results
resOutDir <- file.path(outDir, "Results")
if (!dir.exists(resOutDir)) {
dir.create(resOutDir)
}write.csv(narrowC3ClData, file = file.path(resOutDir, "narrowC3ClChecksRes.csv"))
write.csv(shiftToUnif, file = file.path(resOutDir, "narrowC3ClShiftToUnif.csv"))Look at the C3 clusters
sort(table(c3Clusters), decreasing = TRUE)c3Clusters
CD4 T Cells CD8 T Cells
4607 1562
CD56 Dim NK Cells Naive B Cells
436 315
Classical Monocytes Mucosal-Associated Invariant T Cells
314 276
Gamma-Delta T Cells Nonclassical Monocytes
266 97
CD56 Bright NK Cells IgM Memory B Cells
66 62
Intermediate Monocytes Classical Memory B Cells
49 48
Adaptive NK Cells pDC
24 18
cDC2 Age-associated B Cells
7 6
cDC3 asDC
4 1 largeClNames <- names(which(table(c3Clusters) >= 50))
largeCl <- c3Clusters[(c3Clusters %in% largeClNames), drop = TRUE]
largeCl <- toClustersList(largeCl)lengths(largeCl) CD4 T Cells Gamma-Delta T Cells
4607 266
CD56 Bright NK Cells Mucosal-Associated Invariant T Cells
66 276
CD56 Dim NK Cells Classical Monocytes
436 314
CD8 T Cells Nonclassical Monocytes
1562 97
Naive B Cells IgM Memory B Cells
315 62 resData <- lapply(seq_along(largeCl), FUN = getClData,
objCOTAN = aRunObj, clList = largeCl)
names(resData) <- names(largeCl)
resC3Data <- checkersToDF(resData)
rownames(resC3Data) <- names(largeCl)
resC3Data class
CD4 T Cells SimpleGDIUniformityCheck
Gamma-Delta T Cells SimpleGDIUniformityCheck
CD56 Bright NK Cells SimpleGDIUniformityCheck
Mucosal-Associated Invariant T Cells SimpleGDIUniformityCheck
CD56 Dim NK Cells SimpleGDIUniformityCheck
Classical Monocytes SimpleGDIUniformityCheck
CD8 T Cells SimpleGDIUniformityCheck
Nonclassical Monocytes SimpleGDIUniformityCheck
Naive B Cells SimpleGDIUniformityCheck
IgM Memory B Cells SimpleGDIUniformityCheck
check.isCheckAbove check.GDIThreshold
CD4 T Cells FALSE 1.4
Gamma-Delta T Cells FALSE 1.4
CD56 Bright NK Cells FALSE 1.4
Mucosal-Associated Invariant T Cells FALSE 1.4
CD56 Dim NK Cells FALSE 1.4
Classical Monocytes FALSE 1.4
CD8 T Cells FALSE 1.4
Nonclassical Monocytes FALSE 1.4
Naive B Cells FALSE 1.4
IgM Memory B Cells FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond
CD4 T Cells 0.01 0
Gamma-Delta T Cells 0.01 0
CD56 Bright NK Cells 0.01 0
Mucosal-Associated Invariant T Cells 0.01 0
CD56 Dim NK Cells 0.01 0
Classical Monocytes 0.01 0
CD8 T Cells 0.01 0
Nonclassical Monocytes 0.01 0
Naive B Cells 0.01 0
IgM Memory B Cells 0.01 0
check.fractionBeyond check.thresholdRank
CD4 T Cells 0.59025609 0
Gamma-Delta T Cells 0.13528322 0
CD56 Bright NK Cells 0.01400994 0
Mucosal-Associated Invariant T Cells 0.06778861 0
CD56 Dim NK Cells 0.10958296 0
Classical Monocytes 0.04537929 0
CD8 T Cells 0.40040937 0
Nonclassical Monocytes 0.03663659 0
Naive B Cells 0.17047184 0
IgM Memory B Cells 0.03812241 0
check.quantileAtRatio check.quantileAtRank
CD4 T Cells 2.844100 NaN
Gamma-Delta T Cells 1.630913 NaN
CD56 Bright NK Cells 1.418669 NaN
Mucosal-Associated Invariant T Cells 1.527188 NaN
CD56 Dim NK Cells 1.606543 NaN
Classical Monocytes 1.479715 NaN
CD8 T Cells 2.367379 NaN
Nonclassical Monocytes 1.491915 NaN
Naive B Cells 1.774201 NaN
IgM Memory B Cells 1.504193 NaN
isUniform clusterSize
CD4 T Cells FALSE 4607
Gamma-Delta T Cells FALSE 266
CD56 Bright NK Cells FALSE 66
Mucosal-Associated Invariant T Cells FALSE 276
CD56 Dim NK Cells FALSE 436
Classical Monocytes FALSE 314
CD8 T Cells FALSE 1562
Nonclassical Monocytes FALSE 97
Naive B Cells FALSE 315
IgM Memory B Cells FALSE 62shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`CD4 T Cells`
[1] 1.4441
$`Gamma-Delta T Cells`
[1] 0.2309134
$`CD56 Bright NK Cells`
[1] 0.01866903
$`Mucosal-Associated Invariant T Cells`
[1] 0.1271878
$`CD56 Dim NK Cells`
[1] 0.206543
$`Classical Monocytes`
[1] 0.07971509
$`CD8 T Cells`
[1] 0.9673792
$`Nonclassical Monocytes`
[1] 0.09191463
$`Naive B Cells`
[1] 0.3742006
$`IgM Memory B Cells`
[1] 0.1041927write.csv(resC3Data, file = file.path(resOutDir, "c3ClChecksRes.csv" ))
write.csv(shiftToUnif, file = file.path(resOutDir, "c3ClShiftToUnif.csv"))Coex clustering for CD4 T cells
cellsToDrop <- names(c3Clusters)[c3Clusters != "CD4 T Cells"]
cd4TObj <- dropGenesCells(aRunObj, cells = cellsToDrop)
cd4TObj <- addElementToMetaDataset(
cd4TObj, tag = datasetTags()[["cond"]], value = "CD4 T Cells"
)
cd4TObj <- proceedToCoex(cd4TObj, cores = 3L)fileNameOut <- paste0("CD4Tcells-Run_", thisRun, "-Cleaned", ".RDS")
saveRDS(cd4TObj, file = file.path(outDir, fileNameOut))cd4TObj <- readRDS(file.path(outDir, paste0("CD4Tcells-Run_", thisRun, "-Cleaned", ".RDS")))#table(getClusters(cd4TObj, clName = "Sample_C3"))Create clusterizations
addMultipleClustering <- function(
obj,
numComp = 20L,
range_resolution_min,
range_resolution_max,
step
) {
redDataM <- calculateReducedDataMatrix(
obj, useCoexEigen = TRUE, dataMethod = "LL", numComp = numComp + 15L)
sampleID <- niceFactorLevels(
base::droplevels(getCondition(obj, condName = "Sample.IDs"))
)
samples <- levels(sampleID)
assert_that(length(samples) == 2L)
cells1 <- sampleID == samples[[1L]]
cells2 <- sampleID == samples[[2L]]
# we split the clusters keeping separate the Sample.IDs
FindNeighborsTest1 <-
Seurat::FindNeighbors(redDataM[cells1, seq_len(numComp)])
FindNeighborsTest2 <-
Seurat::FindNeighbors(redDataM[cells2, seq_len(numComp)])
resolutionsList <-
seq(from = range_resolution_min, to = range_resolution_max, by = step)
for(resolution in resolutionsList){
print(resolution)
seuratClusters1 <-
Seurat:::RunModularityClustering(
FindNeighborsTest1$snn,
resolution = resolution,
algorithm = 2L, # Louvain (refined)
random.seed = 137 # controls igraph::cluster_louvain()
)
seuratClusters1 <- paste0(samples[[1L]], "_", seuratClusters1)
names(seuratClusters1) <- getCells(obj)[cells1]
seuratClusters2 <-
Seurat:::RunModularityClustering(
FindNeighborsTest2$snn,
resolution = resolution,
algorithm = 2L, # Louvain (refined)
random.seed = 137 # controls igraph::cluster_louvain()
)
seuratClusters2 <- paste0(samples[[2L]], "_", seuratClusters2)
names(seuratClusters2) <- getCells(obj)[cells2]
seuratClusters <- c(seuratClusters1, seuratClusters2)
seuratClusters <- seuratClusters[getCells(obj)]
print(length(unique(seuratClusters)))
obj <-
addClusterization(
obj,
clName = paste0("LL_New_", resolution),
clusters = seuratClusters,
override = TRUE
)
}
return(obj)
}cd4TObj <-
addMultipleClustering(cd4TObj, numComp = 20L,
range_resolution_min = 0.25,
range_resolution_max = 1.1, step = 0.05)newLLClust <- grep(x = getClusterizations(cd4TObj),
pattern = "^LL_New_", value = TRUE)
numClust <-
vapply(newLLClust,
function(name, obj) {
nlevels(getClusters(obj, clName = name))
}, FUN.VALUE = integer(1), obj = cd4TObj)
names(numClust) <- newLLClust
numClustfileNameOut <- paste0("CD4Tcells-Run_", thisRun, "-Cleaned", ".RDS")
saveRDS(cd4TObj, file = file.path(outDir, fileNameOut))Clusters with resolution 0.3
newClusters <- getClusters(cd4TObj, clName = "LL_New_0.3")
sort(table(newClusters), decreasing = TRUE)newClusters
424_0 095_0 424_1 095_1 095_2 095_3 424_2 424_3
1262 1079 748 742 482 175 95 24 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 095_3 424_0 424_1 424_2
1079 742 482 175 1262 748 95 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd4TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
095_3 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 0.0847773163
095_1 0.01 0 0.0002311782
095_2 0.01 0 0.1416727806
095_3 0.01 0 0.0594585477
424_0 0.01 0 0.3119199006
424_1 0.01 0 0.2398505056
424_2 0.01 0 0.0529509559
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.526399 NaN FALSE
095_1 0 1.342421 NaN TRUE
095_2 0 1.668307 NaN FALSE
095_3 0 1.508794 NaN FALSE
424_0 0 2.099447 NaN FALSE
424_1 0 1.936283 NaN FALSE
424_2 0 1.527378 NaN FALSE
clusterSize
095_0 1079
095_1 742
095_2 482
095_3 175
424_0 1262
424_1 748
424_2 95shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] 0.1263991
$`095_1`
[1] -0.05757905
$`095_2`
[1] 0.2683072
$`095_3`
[1] 0.1087943
$`424_0`
[1] 0.6994471
$`424_1`
[1] 0.536283
$`424_2`
[1] 0.1273782Clusters with resolution 0.35
newClusters <- getClusters(cd4TObj, clName = "LL_New_0.35")
sort(table(newClusters), decreasing = TRUE)newClusters
095_0 095_1 424_0 424_1 424_2 095_2 095_3 424_3 424_4
1070 750 736 728 545 483 175 96 24 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 095_3 424_0 424_1 424_2 424_3
1070 750 483 175 736 728 545 96 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd4TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
095_3 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
424_3 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 0.0822889064
095_1 0.01 0 0.0001537634
095_2 0.01 0 0.1415682535
095_3 0.01 0 0.0594585477
424_0 0.01 0 0.2337738620
424_1 0.01 0 0.2498904310
424_2 0.01 0 0.1516126561
424_3 0.01 0 0.0546330123
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.525003 NaN FALSE
095_1 0 1.342354 NaN TRUE
095_2 0 1.668808 NaN FALSE
095_3 0 1.508794 NaN FALSE
424_0 0 1.925057 NaN FALSE
424_1 0 1.922262 NaN FALSE
424_2 0 1.704905 NaN FALSE
424_3 0 1.530236 NaN FALSE
clusterSize
095_0 1070
095_1 750
095_2 483
095_3 175
424_0 736
424_1 728
424_2 545
424_3 96shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] 0.1250025
$`095_1`
[1] -0.05764559
$`095_2`
[1] 0.2688082
$`095_3`
[1] 0.1087943
$`424_0`
[1] 0.5250574
$`424_1`
[1] 0.5222622
$`424_2`
[1] 0.3049046
$`424_3`
[1] 0.1302359Clusters with resolution 0.4
newClusters <- getClusters(cd4TObj, clName = "LL_New_0.4")
sort(table(newClusters), decreasing = TRUE)newClusters
095_0 095_1 424_0 424_1 095_2 424_2 424_3 095_3 424_4 424_5
1078 742 725 545 483 472 268 175 95 24 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 095_3 424_0 424_1 424_2 424_3 424_4
1078 742 483 175 725 545 472 268 95 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd4TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
095_3 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
424_3 SimpleGDIUniformityCheck FALSE 1.4
424_4 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 0.0847057139
095_1 0.01 0 0.0001540951
095_2 0.01 0 0.1415682535
095_3 0.01 0 0.0594585477
424_0 0.01 0 0.2425373134
424_1 0.01 0 0.1521655807
424_2 0.01 0 0.0635024312
424_3 0.01 0 0.0000000000
424_4 0.01 0 0.0562396007
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.526299 NaN FALSE
095_1 0 1.341580 NaN TRUE
095_2 0 1.668808 NaN FALSE
095_3 0 1.508794 NaN FALSE
424_0 0 1.896355 NaN FALSE
424_1 0 1.705159 NaN FALSE
424_2 0 1.497070 NaN FALSE
424_3 0 1.302151 NaN TRUE
424_4 0 1.532463 NaN FALSE
clusterSize
095_0 1078
095_1 742
095_2 483
095_3 175
424_0 725
424_1 545
424_2 472
424_3 268
424_4 95shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] 0.1262995
$`095_1`
[1] -0.05841967
$`095_2`
[1] 0.2688082
$`095_3`
[1] 0.1087943
$`424_0`
[1] 0.4963551
$`424_1`
[1] 0.3051592
$`424_2`
[1] 0.09707011
$`424_3`
[1] -0.09784929
$`424_4`
[1] 0.1324631Clusters with resolution 0.55
newClusters <- getClusters(cd4TObj, clName = "LL_New_0.55")
sort(table(newClusters), decreasing = TRUE)newClusters
095_0 095_1 424_0 095_2 424_1 095_3 424_2 424_3 424_4 095_4 424_5 424_6
723 664 545 492 468 412 381 342 270 187 99 24 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 095_3 095_4 424_0 424_1 424_2 424_3 424_4 424_5
723 664 492 412 187 545 468 381 342 270 99 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd4TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
095_3 SimpleGDIUniformityCheck FALSE 1.4
095_4 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
424_3 SimpleGDIUniformityCheck FALSE 1.4
424_4 SimpleGDIUniformityCheck FALSE 1.4
424_5 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 0.0045568132
095_1 0.01 0 0.0001569735
095_2 0.01 0 0.1421025603
095_3 0.01 0 0.0004355085
095_4 0.01 0 0.0656674395
424_0 0.01 0 0.1474957880
424_1 0.01 0 0.0622457873
424_2 0.01 0 0.1802884615
424_3 0.01 0 0.1196124800
424_4 0.01 0 0.0000000000
424_5 0.01 0 0.0539803257
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.373409 NaN TRUE
095_1 0 1.337626 NaN TRUE
095_2 0 1.670811 NaN FALSE
095_3 0 1.342705 NaN TRUE
095_4 0 1.519373 NaN FALSE
424_0 0 1.697694 NaN FALSE
424_1 0 1.492416 NaN FALSE
424_2 0 1.791380 NaN FALSE
424_3 0 1.608180 NaN FALSE
424_4 0 1.301712 NaN TRUE
424_5 0 1.527187 NaN FALSE
clusterSize
095_0 723
095_1 664
095_2 492
095_3 412
095_4 187
424_0 545
424_1 468
424_2 381
424_3 342
424_4 270
424_5 99shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] -0.02659079
$`095_1`
[1] -0.06237369
$`095_2`
[1] 0.2708105
$`095_3`
[1] -0.05729523
$`095_4`
[1] 0.1193727
$`424_0`
[1] 0.2976945
$`424_1`
[1] 0.09241603
$`424_2`
[1] 0.3913798
$`424_3`
[1] 0.2081797
$`424_4`
[1] -0.09828777
$`424_5`
[1] 0.1271866Clusters with resolution 0.65
newClusters <- getClusters(cd4TObj, clName = "LL_New_0.65")
sort(table(newClusters), decreasing = TRUE)newClusters
095_0 095_1 095_2 424_0 095_3 424_1 424_2 424_3 424_4 424_5 095_4 424_6 424_7
712 662 494 466 419 376 343 291 271 260 191 98 24 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 095_3 095_4 424_0 424_1 424_2 424_3 424_4 424_5 424_6
712 662 494 419 191 466 376 343 291 271 260 98 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd4TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
095_3 SimpleGDIUniformityCheck FALSE 1.4
095_4 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
424_3 SimpleGDIUniformityCheck FALSE 1.4
424_4 SimpleGDIUniformityCheck FALSE 1.4
424_5 SimpleGDIUniformityCheck FALSE 1.4
424_6 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 0.0039793662
095_1 0.01 0 0.0002356083
095_2 0.01 0 0.1437266261
095_3 0.01 0 0.0004350348
095_4 0.01 0 0.0645719941
424_0 0.01 0 0.0609605464
424_1 0.01 0 0.1803328063
424_2 0.01 0 0.1213092241
424_3 0.01 0 0.0790067720
424_4 0.01 0 0.0000000000
424_5 0.01 0 0.0901460738
424_6 0.01 0 0.0552335210
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.370038 NaN TRUE
095_1 0 1.337344 NaN TRUE
095_2 0 1.672623 NaN FALSE
095_3 0 1.344358 NaN TRUE
095_4 0 1.517239 NaN FALSE
424_0 0 1.490428 NaN FALSE
424_1 0 1.788808 NaN FALSE
424_2 0 1.606448 NaN FALSE
424_3 0 1.567566 NaN FALSE
424_4 0 1.301114 NaN TRUE
424_5 0 1.577562 NaN FALSE
424_6 0 1.529502 NaN FALSE
clusterSize
095_0 712
095_1 662
095_2 494
095_3 419
095_4 191
424_0 466
424_1 376
424_2 343
424_3 291
424_4 271
424_5 260
424_6 98shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] -0.02996186
$`095_1`
[1] -0.06265613
$`095_2`
[1] 0.2726227
$`095_3`
[1] -0.05564167
$`095_4`
[1] 0.1172393
$`424_0`
[1] 0.09042757
$`424_1`
[1] 0.3888081
$`424_2`
[1] 0.2064475
$`424_3`
[1] 0.1675658
$`424_4`
[1] -0.09888584
$`424_5`
[1] 0.1775618
$`424_6`
[1] 0.1295017Clusters with resolution 1
newClusters <- getClusters(cd4TObj, clName = "LL_New_1")
sort(table(newClusters), decreasing = TRUE)newClusters
095_0 095_1 095_2 424_0 095_3 424_1 424_2 424_3 424_4 424_5 095_4 424_6 095_5
687 657 495 429 425 362 358 292 271 261 147 132 67
424_7
24 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 095_3 095_4 095_5 424_0 424_1 424_2 424_3 424_4 424_5 424_6
687 657 495 425 147 67 429 362 358 292 271 261 132 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd4TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
095_3 SimpleGDIUniformityCheck FALSE 1.4
095_4 SimpleGDIUniformityCheck FALSE 1.4
095_5 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
424_3 SimpleGDIUniformityCheck FALSE 1.4
424_4 SimpleGDIUniformityCheck FALSE 1.4
424_5 SimpleGDIUniformityCheck FALSE 1.4
424_6 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 0.0005927243
095_1 0.01 0 0.0002359232
095_2 0.01 0 0.1437059859
095_3 0.01 0 0.0004346566
095_4 0.01 0 0.0154902709
095_5 0.01 0 0.0108495931
424_0 0.01 0 0.0561542409
424_1 0.01 0 0.1247251497
424_2 0.01 0 0.1702690447
424_3 0.01 0 0.0833074051
424_4 0.01 0 0.0000000000
424_5 0.01 0 0.0904469905
424_6 0.01 0 0.0533740889
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.345552 NaN TRUE
095_1 0 1.338326 NaN TRUE
095_2 0 1.667833 NaN FALSE
095_3 0 1.342436 NaN TRUE
095_4 0 1.421659 NaN FALSE
095_5 0 1.404116 NaN FALSE
424_0 0 1.490805 NaN FALSE
424_1 0 1.620600 NaN FALSE
424_2 0 1.749649 NaN FALSE
424_3 0 1.578602 NaN FALSE
424_4 0 1.301114 NaN TRUE
424_5 0 1.581919 NaN FALSE
424_6 0 1.511813 NaN FALSE
clusterSize
095_0 687
095_1 657
095_2 495
095_3 425
095_4 147
095_5 67
424_0 429
424_1 362
424_2 358
424_3 292
424_4 271
424_5 261
424_6 132shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] -0.05444768
$`095_1`
[1] -0.06167428
$`095_2`
[1] 0.2678332
$`095_3`
[1] -0.05756446
$`095_4`
[1] 0.02165941
$`095_5`
[1] 0.004116039
$`424_0`
[1] 0.09080506
$`424_1`
[1] 0.2205995
$`424_2`
[1] 0.3496488
$`424_3`
[1] 0.1786015
$`424_4`
[1] -0.09888584
$`424_5`
[1] 0.181919
$`424_6`
[1] 0.1118133Clusters with resolution 1.05
newClusters <- getClusters(cd4TObj, clName = "LL_New_1.05")
sort(table(newClusters), decreasing = TRUE)newClusters
095_0 095_1 095_2 095_3 424_0 424_1 424_2 424_3 424_4 424_5 424_6 095_4 424_7
686 656 496 425 422 340 271 254 251 228 200 148 139
095_5 424_8
67 24 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 095_3 095_4 095_5 424_0 424_1 424_2 424_3 424_4 424_5 424_6
686 656 496 425 148 67 422 340 271 254 251 228 200
424_7
139 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd4TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
095_3 SimpleGDIUniformityCheck FALSE 1.4
095_4 SimpleGDIUniformityCheck FALSE 1.4
095_5 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
424_3 SimpleGDIUniformityCheck FALSE 1.4
424_4 SimpleGDIUniformityCheck FALSE 1.4
424_5 SimpleGDIUniformityCheck FALSE 1.4
424_6 SimpleGDIUniformityCheck FALSE 1.4
424_7 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 5.187106e-04
095_1 0.01 0 1.572822e-04
095_2 0.01 0 1.436533e-01
095_3 0.01 0 4.346566e-04
095_4 0.01 0 1.570840e-02
095_5 0.01 0 1.084959e-02
424_0 0.01 0 5.040411e-02
424_1 0.01 0 1.142487e-01
424_2 0.01 0 0.000000e+00
424_3 0.01 0 7.071689e-02
424_4 0.01 0 9.006085e-03
424_5 0.01 0 8.515711e-05
424_6 0.01 0 4.086353e-03
424_7 0.01 0 5.526152e-02
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.346127 NaN TRUE
095_1 0 1.339001 NaN TRUE
095_2 0 1.668680 NaN FALSE
095_3 0 1.342436 NaN TRUE
095_4 0 1.422347 NaN FALSE
095_5 0 1.404116 NaN FALSE
424_0 0 1.483614 NaN FALSE
424_1 0 1.593147 NaN FALSE
424_2 0 1.301114 NaN TRUE
424_3 0 1.543700 NaN FALSE
424_4 0 1.396705 NaN TRUE
424_5 0 1.329809 NaN TRUE
424_6 0 1.372698 NaN TRUE
424_7 0 1.517190 NaN FALSE
clusterSize
095_0 686
095_1 656
095_2 496
095_3 425
095_4 148
095_5 67
424_0 422
424_1 340
424_2 271
424_3 254
424_4 251
424_5 228
424_6 200
424_7 139shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] -0.05387256
$`095_1`
[1] -0.06099862
$`095_2`
[1] 0.2686802
$`095_3`
[1] -0.05756446
$`095_4`
[1] 0.02234719
$`095_5`
[1] 0.004116039
$`424_0`
[1] 0.08361447
$`424_1`
[1] 0.1931467
$`424_2`
[1] -0.09888584
$`424_3`
[1] 0.1436996
$`424_4`
[1] -0.003295144
$`424_5`
[1] -0.07019073
$`424_6`
[1] -0.02730181
$`424_7`
[1] 0.1171897Clusters with resolution 1.1
newClusters <- getClusters(cd4TObj, clName = "LL_New_1.1")
sort(table(newClusters), decreasing = TRUE)newClusters
095_0 095_1 095_2 095_3 424_0 424_1 424_2 424_3 424_4 424_5 424_6 095_4 424_7
681 654 496 428 421 347 271 253 241 230 201 151 103
095_5 424_8 424_9
68 38 24 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 095_3 095_4 095_5 424_0 424_1 424_2 424_3 424_4 424_5 424_6
681 654 496 428 151 68 421 347 271 253 241 230 201
424_7
103 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd4TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
095_3 SimpleGDIUniformityCheck FALSE 1.4
095_4 SimpleGDIUniformityCheck FALSE 1.4
095_5 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
424_3 SimpleGDIUniformityCheck FALSE 1.4
424_4 SimpleGDIUniformityCheck FALSE 1.4
424_5 SimpleGDIUniformityCheck FALSE 1.4
424_6 SimpleGDIUniformityCheck FALSE 1.4
424_7 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 7.423905e-05
095_1 0.01 0 1.573812e-04
095_2 0.01 0 1.436533e-01
095_3 0.01 0 5.070259e-04
095_4 0.01 0 1.545371e-02
095_5 0.01 0 1.933201e-02
424_0 0.01 0 5.055850e-02
424_1 0.01 0 1.185681e-01
424_2 0.01 0 0.000000e+00
424_3 0.01 0 7.044994e-02
424_4 0.01 0 5.404078e-03
424_5 0.01 0 1.695778e-04
424_6 0.01 0 3.004391e-03
424_7 0.01 0 5.594750e-02
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.335691 NaN TRUE
095_1 0 1.339208 NaN TRUE
095_2 0 1.668680 NaN FALSE
095_3 0 1.343202 NaN TRUE
095_4 0 1.423718 NaN FALSE
095_5 0 1.432058 NaN FALSE
424_0 0 1.482916 NaN FALSE
424_1 0 1.597820 NaN FALSE
424_2 0 1.301114 NaN TRUE
424_3 0 1.542328 NaN FALSE
424_4 0 1.381597 NaN TRUE
424_5 0 1.332733 NaN TRUE
424_6 0 1.368131 NaN TRUE
424_7 0 1.525626 NaN FALSE
clusterSize
095_0 681
095_1 654
095_2 496
095_3 428
095_4 151
095_5 68
424_0 421
424_1 347
424_2 271
424_3 253
424_4 241
424_5 230
424_6 201
424_7 103shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] -0.06430948
$`095_1`
[1] -0.06079204
$`095_2`
[1] 0.2686802
$`095_3`
[1] -0.05679758
$`095_4`
[1] 0.02371812
$`095_5`
[1] 0.0320576
$`424_0`
[1] 0.08291566
$`424_1`
[1] 0.1978204
$`424_2`
[1] -0.09888584
$`424_3`
[1] 0.1423277
$`424_4`
[1] -0.01840275
$`424_5`
[1] -0.06726691
$`424_6`
[1] -0.0318691
$`424_7`
[1] 0.1256262Coex clustering for CD8 T cells
cellsToDrop <- names(c3Clusters)[c3Clusters != "CD8 T Cells"]
cd8TObj <- dropGenesCells(aRunObj, cells = cellsToDrop)
cd8TObj <- addElementToMetaDataset(
cd8TObj, tag = datasetTags()[["cond"]], value = "CD8 T Cells"
)
cd8TObj <- proceedToCoex(cd8TObj, cores = 3L)fileNameOut <- paste0("CD8Tcells-Run_", thisRun, "-Cleaned", ".RDS")
saveRDS(cd8TObj, file = file.path(outDir, fileNameOut))cd8TObj <- readRDS(file.path(outDir, paste0("CD8Tcells-Run_", thisRun, "-Cleaned", ".RDS")))#table(getClusters(cd8TObj, clName = "Sample_C3"))cd8TObj <-
addMultipleClustering(cd8TObj, numComp = 20L,
range_resolution_min = 0.25,
range_resolution_max = 1.1, step = 0.05)newLLClust <- grep(x = getClusterizations(cd8TObj),
pattern = "^LL_New_", value = TRUE)
numClust <-
vapply(newLLClust,
function(name, obj) {
nlevels(getClusters(obj, clName = name))
}, FUN.VALUE = integer(1), obj = cd8TObj)
names(numClust) <- newLLClust
numClustfileNameOut <- paste0("CD8Tcells-Run_", thisRun, "-Cleaned", ".RDS")
saveRDS(cd8TObj, file = file.path(outDir, fileNameOut))Clusters with resolution 0.75
newClusters <- getClusters(cd8TObj, clName = "LL_New_0.75")
sort(table(newClusters), decreasing = TRUE)newClusters
424_0 424_1 095_0 095_1 424_2 095_2
318 305 297 283 222 137 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 424_0 424_1 424_2
297 283 137 318 305 222 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd8TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 8.149959e-05
095_1 0.01 0 8.865790e-03
095_2 0.01 0 8.618663e-03
424_0 0.01 0 1.034749e-01
424_1 0.01 0 1.401810e-01
424_2 0.01 0 7.426345e-02
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.317952 NaN TRUE
095_1 0 1.395932 NaN TRUE
095_2 0 1.392219 NaN TRUE
424_0 0 1.603377 NaN FALSE
424_1 0 1.665812 NaN FALSE
424_2 0 1.547176 NaN FALSE
clusterSize
095_0 297
095_1 283
095_2 137
424_0 318
424_1 305
424_2 222shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] -0.08204785
$`095_1`
[1] -0.004068082
$`095_2`
[1] -0.007781029
$`424_0`
[1] 0.2033772
$`424_1`
[1] 0.2658123
$`424_2`
[1] 0.1471758Clusters with resolution 0.9
newClusters <- getClusters(cd8TObj, clName = "LL_New_0.9")
sort(table(newClusters), decreasing = TRUE)newClusters
095_0 424_0 095_1 424_1 424_2 424_3 424_4 095_2 095_3
267 213 182 169 166 152 145 137 131 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 095_3 424_0 424_1 424_2 424_3 424_4
267 182 137 131 213 169 166 152 145 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd8TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
095_3 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
424_3 SimpleGDIUniformityCheck FALSE 1.4
424_4 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 0.0072975880
095_1 0.01 0 0.0000000000
095_2 0.01 0 0.0086186633
095_3 0.01 0 0.0000000000
424_0 0.01 0 0.0617233486
424_1 0.01 0 0.0059409254
424_2 0.01 0 0.0013562387
424_3 0.01 0 0.0001601666
424_4 0.01 0 0.0794032724
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.391408 NaN TRUE
095_1 0 1.297215 NaN TRUE
095_2 0 1.392219 NaN TRUE
095_3 0 1.235970 NaN TRUE
424_0 0 1.529457 NaN FALSE
424_1 0 1.383551 NaN TRUE
424_2 0 1.349994 NaN TRUE
424_3 0 1.308943 NaN TRUE
424_4 0 1.603209 NaN FALSE
clusterSize
095_0 267
095_1 182
095_2 137
095_3 131
424_0 213
424_1 169
424_2 166
424_3 152
424_4 145shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] -0.008591753
$`095_1`
[1] -0.1027848
$`095_2`
[1] -0.007781029
$`095_3`
[1] -0.1640298
$`424_0`
[1] 0.1294572
$`424_1`
[1] -0.01644878
$`424_2`
[1] -0.05000604
$`424_3`
[1] -0.09105709
$`424_4`
[1] 0.2032093Clusters with resolution 1.1
newClusters <- getClusters(cd8TObj, clName = "LL_New_1.1")
sort(table(newClusters), decreasing = TRUE)newClusters
095_0 424_0 095_1 424_1 424_2 424_3 095_2 095_3 424_4 424_5
267 212 180 167 166 152 137 133 101 47 largeClNames <- names(which(table(newClusters) >= 50))
largeNewCl <- newClusters[newClusters %in% largeClNames]
largeNewCl <- toClustersList(largeNewCl)lengths(largeNewCl)095_0 095_1 095_2 095_3 424_0 424_1 424_2 424_3 424_4
267 180 137 133 212 167 166 152 101 resData <- lapply(seq_along(largeNewCl), FUN = getClData,
objCOTAN = cd8TObj, clList = largeNewCl)
names(resData) <- names(largeNewCl)
resNewData <- checkersToDF(resData)
rownames(resNewData) <- names(largeNewCl)
resNewData class check.isCheckAbove check.GDIThreshold
095_0 SimpleGDIUniformityCheck FALSE 1.4
095_1 SimpleGDIUniformityCheck FALSE 1.4
095_2 SimpleGDIUniformityCheck FALSE 1.4
095_3 SimpleGDIUniformityCheck FALSE 1.4
424_0 SimpleGDIUniformityCheck FALSE 1.4
424_1 SimpleGDIUniformityCheck FALSE 1.4
424_2 SimpleGDIUniformityCheck FALSE 1.4
424_3 SimpleGDIUniformityCheck FALSE 1.4
424_4 SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
095_0 0.01 0 0.0072975880
095_1 0.01 0 0.0000000000
095_2 0.01 0 0.0086186633
095_3 0.01 0 0.0000000000
424_0 0.01 0 0.0623567921
424_1 0.01 0 0.0073745077
424_2 0.01 0 0.0013562387
424_3 0.01 0 0.0001601666
424_4 0.01 0 0.0000000000
check.thresholdRank check.quantileAtRatio check.quantileAtRank isUniform
095_0 0 1.391408 NaN TRUE
095_1 0 1.297197 NaN TRUE
095_2 0 1.392219 NaN TRUE
095_3 0 1.235968 NaN TRUE
424_0 0 1.529741 NaN FALSE
424_1 0 1.387691 NaN TRUE
424_2 0 1.349994 NaN TRUE
424_3 0 1.308943 NaN TRUE
424_4 0 1.255680 NaN TRUE
clusterSize
095_0 267
095_1 180
095_2 137
095_3 133
424_0 212
424_1 167
424_2 166
424_3 152
424_4 101shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$`095_0`
[1] -0.008591753
$`095_1`
[1] -0.1028025
$`095_2`
[1] -0.007781029
$`095_3`
[1] -0.164032
$`424_0`
[1] 0.1297409
$`424_1`
[1] -0.01230937
$`424_2`
[1] -0.05000604
$`424_3`
[1] -0.09105709
$`424_4`
[1] -0.1443197Relevant clusters
cd4Clusters <- toClustersList(getClusters(cd4TObj, clName = "LL_New_1.05"))
cd8Clusters <- toClustersList(getClusters(cd8TObj, clName = "LL_New_0.9"))
relCD4ClNames <- c("095_0", "095_1", "095_3", "095_5",
"424_2", "424_4", "424_5", "424_6")
relCD8ClNames <- c("095_0", "095_1", "095_2", "095_3",
"424_1", "424_2", "424_3")
relCD4Cl <- cd4Clusters[relCD4ClNames]
relCD8Cl <- cd8Clusters[relCD8ClNames]
relevantClustersList <- append(relCD4Cl, relCD8Cl)
names(relevantClustersList) <-
c(paste0("CD4_", names(relCD4Cl)), paste0("CD8_", names(relCD8Cl)))
otherCells <- setdiff(getCells(aRunObj), unlist(relevantClustersList))
assert_that(length(otherCells) + sum(lengths(relevantClustersList)) == getNumCells(aRunObj))[1] TRUErelevantClustersList <- append(relevantClustersList, list(Other = otherCells))
assert_that(sum(lengths(relevantClustersList)) == getNumCells(aRunObj))[1] TRUErelevantClusters <- fromClustersList(relevantClustersList)
aRunObj <-
addClusterization(aRunObj, clName = "SelUnifCl", override = TRUE,
clusters = relevantClusters[getCells(aRunObj)])Save COTAN obj
fileNameOut <- paste0(globalCondition, "-Run_", thisRun, "-Cleaned", ".RDS")
saveRDS(aRunObj, file = file.path(outDir, fileNameOut))Check relevant clusters
sort(table(relevantClusters), decreasing = TRUE)relevantClusters
Other CD4_095_0 CD4_095_1 CD4_095_3 CD4_424_2 CD8_095_0 CD4_424_4 CD4_424_5
4170 686 656 425 271 267 251 228
CD4_424_6 CD8_095_1 CD8_424_1 CD8_424_2 CD8_424_3 CD8_095_2 CD8_095_3 CD4_095_5
200 182 169 166 152 137 131 67 largeClNames <- names(which(table(relevantClusters) >= 50))
largeRelCl <- relevantClusters[relevantClusters %in% largeClNames]
largeRelCl <- toClustersList(largeRelCl)lengths(largeRelCl)CD4_095_0 CD4_095_1 CD4_095_3 CD4_095_5 CD4_424_2 CD4_424_4 CD4_424_5 CD4_424_6
686 656 425 67 271 251 228 200
CD8_095_0 CD8_095_1 CD8_095_2 CD8_095_3 CD8_424_1 CD8_424_2 CD8_424_3 Other
267 182 137 131 169 166 152 4170 resData <- lapply(seq_along(largeRelCl), FUN = getClData,
objCOTAN = aRunObj, clList = largeRelCl)
names(resData) <- names(largeRelCl)
resRelData <- checkersToDF(resData)
rownames(resRelData) <- names(largeRelCl)
resRelData class check.isCheckAbove check.GDIThreshold
CD4_095_0 SimpleGDIUniformityCheck FALSE 1.4
CD4_095_1 SimpleGDIUniformityCheck FALSE 1.4
CD4_095_3 SimpleGDIUniformityCheck FALSE 1.4
CD4_095_5 SimpleGDIUniformityCheck FALSE 1.4
CD4_424_2 SimpleGDIUniformityCheck FALSE 1.4
CD4_424_4 SimpleGDIUniformityCheck FALSE 1.4
CD4_424_5 SimpleGDIUniformityCheck FALSE 1.4
CD4_424_6 SimpleGDIUniformityCheck FALSE 1.4
CD8_095_0 SimpleGDIUniformityCheck FALSE 1.4
CD8_095_1 SimpleGDIUniformityCheck FALSE 1.4
CD8_095_2 SimpleGDIUniformityCheck FALSE 1.4
CD8_095_3 SimpleGDIUniformityCheck FALSE 1.4
CD8_424_1 SimpleGDIUniformityCheck FALSE 1.4
CD8_424_2 SimpleGDIUniformityCheck FALSE 1.4
CD8_424_3 SimpleGDIUniformityCheck FALSE 1.4
Other SimpleGDIUniformityCheck FALSE 1.4
check.maxRatioBeyond check.maxRankBeyond check.fractionBeyond
CD4_095_0 0.01 0 0.0003666496
CD4_095_1 0.01 0 0.0001564945
CD4_095_3 0.01 0 0.0004255017
CD4_095_5 0.01 0 0.0100380755
CD4_424_2 0.01 0 0.0000000000
CD4_424_4 0.01 0 0.0083776382
CD4_424_5 0.01 0 0.0000847745
CD4_424_6 0.01 0 0.0033549371
CD8_095_0 0.01 0 0.0064413459
CD8_095_1 0.01 0 0.0000000000
CD8_095_2 0.01 0 0.0067329763
CD8_095_3 0.01 0 0.0000000000
CD8_424_1 0.01 0 0.0053964301
CD8_424_2 0.01 0 0.0010976948
CD8_424_3 0.01 0 0.0001572203
Other 0.01 0 0.7924281129
check.thresholdRank check.quantileAtRatio check.quantileAtRank
CD4_095_0 0 1.344230 NaN
CD4_095_1 0 1.338319 NaN
CD4_095_3 0 1.337946 NaN
CD4_095_5 0 1.400001 NaN
CD4_424_2 0 1.300256 NaN
CD4_424_4 0 1.394160 NaN
CD4_424_5 0 1.328158 NaN
CD4_424_6 0 1.369559 NaN
CD8_095_0 0 1.388063 NaN
CD8_095_1 0 1.296363 NaN
CD8_095_2 0 1.383290 NaN
CD8_095_3 0 1.231201 NaN
CD8_424_1 0 1.381444 NaN
CD8_424_2 0 1.341080 NaN
CD8_424_3 0 1.302024 NaN
Other 0 4.891091 NaN
isUniform clusterSize
CD4_095_0 TRUE 686
CD4_095_1 TRUE 656
CD4_095_3 TRUE 425
CD4_095_5 FALSE 67
CD4_424_2 TRUE 271
CD4_424_4 TRUE 251
CD4_424_5 TRUE 228
CD4_424_6 TRUE 200
CD8_095_0 TRUE 267
CD8_095_1 TRUE 182
CD8_095_2 TRUE 137
CD8_095_3 TRUE 131
CD8_424_1 TRUE 169
CD8_424_2 TRUE 166
CD8_424_3 TRUE 152
Other FALSE 4170shiftToUnif <- lapply(resData, calculateThresholdShiftToUniformity)
shiftToUnif$CD4_095_0
[1] -0.05577031
$CD4_095_1
[1] -0.06168086
$CD4_095_3
[1] -0.06205365
$CD4_095_5
[1] 6.423258e-07
$CD4_424_2
[1] -0.09974366
$CD4_424_4
[1] -0.005839663
$CD4_424_5
[1] -0.07184202
$CD4_424_6
[1] -0.03044132
$CD8_095_0
[1] -0.01193724
$CD8_095_1
[1] -0.1036367
$CD8_095_2
[1] -0.01670988
$CD8_095_3
[1] -0.1687987
$CD8_424_1
[1] -0.01855575
$CD8_424_2
[1] -0.05892038
$CD8_424_3
[1] -0.09797634
$Other
[1] 3.491091write.csv(resRelData, file = file.path(resOutDir, "relevantClChecksRes.csv"))Session info
Sys.time()[1] "2026-02-06 07:04:47 CET"#Sys.info()
sessionInfo()R version 4.5.2 (2025-10-31)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 22.04.5 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.10.0
LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.10.0 LAPACK version 3.10.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: Europe/Rome
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] COTAN_2.11.1 Matrix_1.7-4 conflicted_1.2.0 zeallot_0.2.0
[5] ggplot2_4.0.1 assertthat_0.2.1
loaded via a namespace (and not attached):
[1] RcppAnnoy_0.0.22 splines_4.5.2
[3] later_1.4.2 tibble_3.3.0
[5] polyclip_1.10-7 fastDummies_1.7.5
[7] lifecycle_1.0.4 doParallel_1.0.17
[9] processx_3.8.6 globals_0.18.0
[11] lattice_0.22-7 MASS_7.3-65
[13] ggdist_3.3.3 dendextend_1.19.0
[15] magrittr_2.0.4 plotly_4.12.0
[17] rmarkdown_2.29 yaml_2.3.12
[19] httpuv_1.6.16 otel_0.2.0
[21] Seurat_5.4.0 sctransform_0.4.2
[23] spam_2.11-1 sp_2.2-0
[25] spatstat.sparse_3.1-0 reticulate_1.44.1
[27] cowplot_1.2.0 pbapply_1.7-2
[29] RColorBrewer_1.1-3 abind_1.4-8
[31] Rtsne_0.17 GenomicRanges_1.62.1
[33] purrr_1.2.0 BiocGenerics_0.56.0
[35] coro_1.1.0 torch_0.16.3
[37] circlize_0.4.16 GenomeInfoDbData_1.2.14
[39] IRanges_2.44.0 S4Vectors_0.48.0
[41] ggrepel_0.9.6 irlba_2.3.5.1
[43] listenv_0.10.0 spatstat.utils_3.2-1
[45] goftest_1.2-3 RSpectra_0.16-2
[47] spatstat.random_3.4-3 fitdistrplus_1.2-2
[49] parallelly_1.46.0 codetools_0.2-20
[51] DelayedArray_0.36.0 tidyselect_1.2.1
[53] shape_1.4.6.1 UCSC.utils_1.4.0
[55] farver_2.1.2 ScaledMatrix_1.16.0
[57] viridis_0.6.5 matrixStats_1.5.0
[59] stats4_4.5.2 spatstat.explore_3.7-0
[61] Seqinfo_1.0.0 jsonlite_2.0.0
[63] GetoptLong_1.1.0 progressr_0.18.0
[65] ggridges_0.5.6 survival_3.8-3
[67] iterators_1.0.14 foreach_1.5.2
[69] tools_4.5.2 ica_1.0-3
[71] Rcpp_1.1.0 glue_1.8.0
[73] gridExtra_2.3 SparseArray_1.10.8
[75] xfun_0.52 MatrixGenerics_1.22.0
[77] distributional_0.6.0 ggthemes_5.2.0
[79] GenomeInfoDb_1.44.0 dplyr_1.1.4
[81] withr_3.0.2 fastmap_1.2.0
[83] callr_3.7.6 digest_0.6.37
[85] rsvd_1.0.5 parallelDist_0.2.6
[87] R6_2.6.1 mime_0.13
[89] colorspace_2.1-1 scattermore_1.2
[91] tensor_1.5 spatstat.data_3.1-9
[93] tidyr_1.3.1 generics_0.1.3
[95] data.table_1.18.0 httr_1.4.7
[97] htmlwidgets_1.6.4 S4Arrays_1.10.1
[99] uwot_0.2.3 pkgconfig_2.0.3
[101] gtable_0.3.6 ComplexHeatmap_2.26.0
[103] lmtest_0.9-40 S7_0.2.1
[105] SingleCellExperiment_1.32.0 XVector_0.50.0
[107] htmltools_0.5.8.1 dotCall64_1.2
[109] zigg_0.0.2 clue_0.3-66
[111] SeuratObject_5.3.0 scales_1.4.0
[113] Biobase_2.70.0 png_0.1-8
[115] spatstat.univar_3.1-6 knitr_1.50
[117] reshape2_1.4.4 rjson_0.2.23
[119] nlme_3.1-168 proxy_0.4-29
[121] cachem_1.1.0 zoo_1.8-14
[123] GlobalOptions_0.1.2 stringr_1.6.0
[125] KernSmooth_2.23-26 parallel_4.5.2
[127] miniUI_0.1.2 pillar_1.11.1
[129] grid_4.5.2 vctrs_0.7.0
[131] RANN_2.6.2 promises_1.5.0
[133] BiocSingular_1.26.1 beachmat_2.26.0
[135] xtable_1.8-4 cluster_2.1.8.1
[137] evaluate_1.0.5 cli_3.6.5
[139] compiler_4.5.2 rlang_1.1.7
[141] crayon_1.5.3 future.apply_1.20.0
[143] labeling_0.4.3 ps_1.9.1
[145] plyr_1.8.9 stringi_1.8.7
[147] viridisLite_0.4.2 deldir_2.0-4
[149] BiocParallel_1.44.0 lazyeval_0.2.2
[151] spatstat.geom_3.7-0 RcppHNSW_0.6.0
[153] patchwork_1.3.2 bit64_4.6.0-1
[155] future_1.69.0 shiny_1.12.1
[157] SummarizedExperiment_1.38.1 ROCR_1.0-11
[159] Rfast_2.1.5.1 igraph_2.2.1
[161] memoise_2.0.1 RcppParallel_5.1.10
[163] bit_4.6.0