# Util libs
library(assertthat)
library(ggplot2)
library(zeallot)
library(conflicted)
library(Matrix)
library(stringr)
library(tidyverse)
library(ComplexHeatmap)
# Data processing libs
if (!suppressWarnings(require(COTAN))) {
devtools::load_all("~/dev/COTAN/COTAN/")
}
conflicts_prefer(zeallot::`%->%`, zeallot::`%<-%`)
options(parallelly.fork.enable = TRUE)
setLoggingLevel(2L)Relevant Clusters Distance Evaluation using Run 40
Preamble
inDir <- file.path("Data/Brown_PBMC_Datasets/")
outDir <- file.path("Results/GDI_Sensitivity/PBMCBrownRun40/")
list.files(path = inDir, pattern = "\\.RDS$") [1] "capillary_blood_samples_pbmcs-Run_28.RDS"
[2] "capillary_blood_samples_pbmcs-Run_40-Cleaned.RDS"
[3] "capillary_blood_samples_pbmcs-Run_40.RDS"
[4] "capillary_blood_samples_pbmcs-Run_41-Cleaned.RDS"
[5] "capillary_blood_samples_pbmcs-Run_41.RDS"
[6] "capillary_blood_samples_pbmcs-Run_62.RDS"
[7] "capillary_blood_samples_pbmcs-Run_77-Cleaned.RDS"
[8] "capillary_blood_samples_pbmcs-Run_77.RDS"
[9] "capillary_blood_samples_pbmcs.RDS"
[10] "CD4Tcells-Run_40-Cleaned.RDS"
[11] "Run_40Cl_0CD4TcellsRawData.RDS"
[12] "Run_40Cl_1CD4TcellsRawData.RDS"
[13] "Run_40Cl_5CD4TcellsRawData.RDS"
[14] "Run_40Cl_7CD4TcellsRawData.RDS"
[15] "Run_40Cl_8CD4TcellsRawData.RDS"
[16] "Run_40Cl_9CD4TcellsRawData.RDS" globalCondition <- "capillary_blood_samples_pbmcs"
thisRun <- "40"
fileNameIn <- paste0(globalCondition, "-Run_", thisRun, "-Cleaned.RDS")
setLoggingFile(file.path(inDir, paste0("ClustersDistance_Run", thisRun, ".log")))Load COTAN object
aRunObj <- readRDS(file = file.path(inDir, fileNameIn))
sampleCondition <- getMetadataElement(aRunObj, datasetTags()[["cond"]])[[1L]]
getClusterizations(aRunObj) [1] "cell_type_level_1" "cell_type_level_2" "cell_type_level_3"
[4] "cell_type_level_4" "c1" "c2"
[7] "c3" "c4" "Sample_C3"
[10] "SelUnifCl" relevantClusters <- getClusters(aRunObj, clName = "SelUnifCl")relevantClustersList <- toClustersList(relevantClusters)
relevantClustersList <- relevantClustersList[names(relevantClustersList) != "Other"]Defining the two distances
To roughly determine the cluster distances we decided to test two simple Euclidean distances:
over the mean of the 0/1 raw matrix
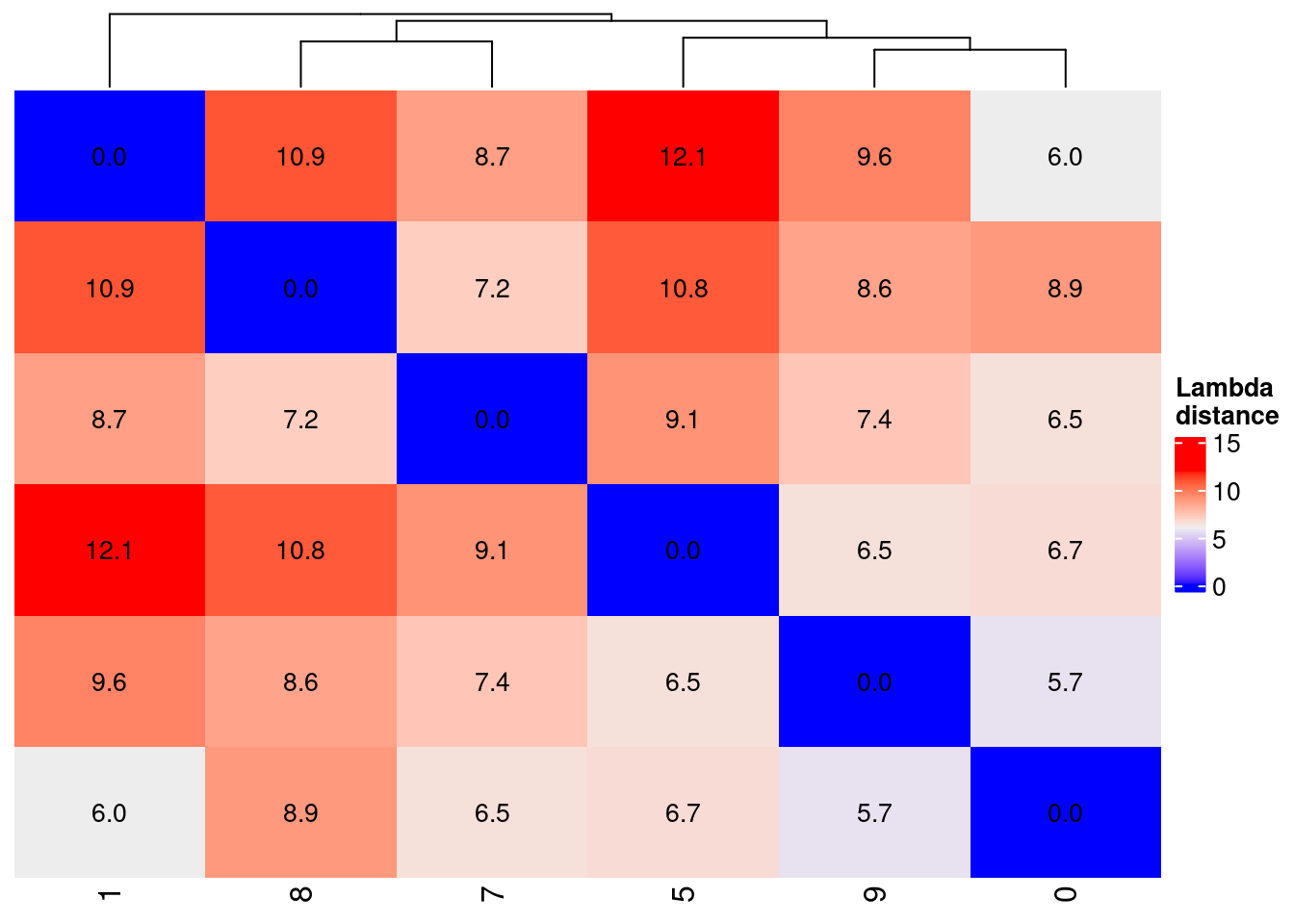
over the \(1-e^{-\lambda}\) where \(\lambda\) is the average expression for the genes.
allZeroOne <- data.frame(row.names = getGenes(aRunObj))
allProbOne <- data.frame(row.names = getGenes(aRunObj))
for (clName in names(relevantClustersList)) {
print(paste("Calculating statistics for custer:", clName))
cellsToDrop <- getCells(aRunObj)[relevantClusters != clName]
obj <- dropGenesCells(aRunObj, cells = cellsToDrop)
# for each gene get average presence in a cell
ZeroOne <- rowMeans(getZeroOneProj(obj))
allZeroOne <- cbind(allZeroOne, ZeroOne)
colnames(allZeroOne)[ncol(allZeroOne)] <- clName
# for each gene get the average probability of presence in a cell
obj <- estimateLambdaLinear(obj)
ProbOne <- 1 - exp(-getLambda(obj))
allProbOne <- cbind(allProbOne, ProbOne)
colnames(allProbOne)[ncol(allProbOne)] <- clName
}
head(allZeroOne)
head(allProbOne)Saving the df
saveRDS(allZeroOne, file.path(outDir, "allZeroOne.RDS"))
saveRDS(allProbOne, file.path(outDir, "allProbOne.RDS"))allZeroOne <- readRDS(file.path(outDir, "allZeroOne.RDS"))
allProbOne <- readRDS(file.path(outDir, "allProbOne.RDS"))ProbOne distance
distanceProbOne <-
as.matrix(dist(t(allProbOne), method = "euclidean",
diag = TRUE, upper = TRUE))
saveRDS(distanceProbOne, file.path(outDir, "distanceProbOne.RDS"))
Heatmap(
distanceProbOne,
name = "Lambda\ndistance",
cell_fun = function(j, i, x, y, width, height, fill)
{
grid.text(sprintf("%.1f", distanceProbOne[i, j]), x, y, gp = gpar(fontsize = 10))
},
show_row_dend = FALSE,
show_row_names = FALSE
)
ZeroOne distance
distanceZeroOne <-
as.matrix(dist(t(allZeroOne), method = "euclidean",
diag = TRUE, upper = TRUE))
saveRDS(distanceZeroOne, file.path(outDir, "distanceZeroOne.RDS"))
HeatmapZeroOne <-
Heatmap(
distanceZeroOne,
name = "ZeroOne\ndistance",
cell_fun = function(j, i, x, y, width, height, fill)
{
grid.text(sprintf("%.1f", distanceZeroOne[i, j]),
x, y, gp = gpar(fontsize = 10))
},
show_row_dend = FALSE,
show_row_names = FALSE
)
pdf(file = file.path(outDir,"HeatmapZeroOne.pdf"), width = 6, height = 5)
plot(HeatmapZeroOne)
dev.off()png
2 plot(HeatmapZeroOne)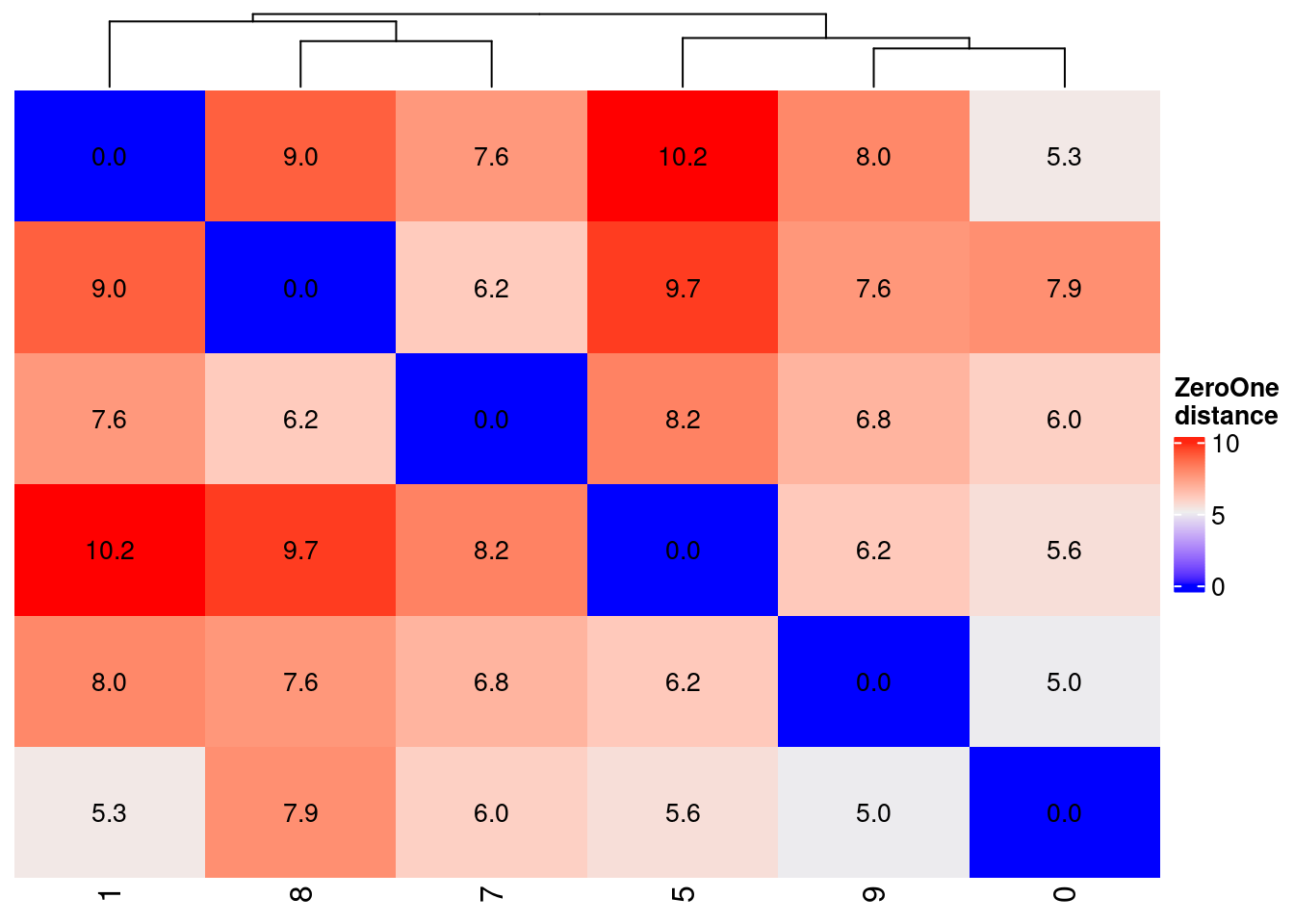
Sys.time()[1] "2026-02-02 15:43:46 CET"#Sys.info()
sessionInfo()R version 4.5.2 (2025-10-31)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 22.04.5 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.10.0
LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.10.0 LAPACK version 3.10.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: Europe/Rome
tzcode source: system (glibc)
attached base packages:
[1] grid stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] COTAN_2.11.1 ComplexHeatmap_2.26.0 lubridate_1.9.4
[4] forcats_1.0.0 dplyr_1.1.4 purrr_1.2.0
[7] readr_2.1.5 tidyr_1.3.1 tibble_3.3.0
[10] tidyverse_2.0.0 stringr_1.6.0 Matrix_1.7-4
[13] conflicted_1.2.0 zeallot_0.2.0 ggplot2_4.0.1
[16] assertthat_0.2.1
loaded via a namespace (and not attached):
[1] RcppAnnoy_0.0.22 splines_4.5.2
[3] later_1.4.2 polyclip_1.10-7
[5] fastDummies_1.7.5 lifecycle_1.0.4
[7] doParallel_1.0.17 globals_0.18.0
[9] lattice_0.22-7 MASS_7.3-65
[11] ggdist_3.3.3 dendextend_1.19.0
[13] magrittr_2.0.4 plotly_4.12.0
[15] rmarkdown_2.29 yaml_2.3.12
[17] httpuv_1.6.16 otel_0.2.0
[19] Seurat_5.4.0 sctransform_0.4.2
[21] spam_2.11-1 sp_2.2-0
[23] spatstat.sparse_3.1-0 reticulate_1.44.1
[25] cowplot_1.2.0 pbapply_1.7-2
[27] RColorBrewer_1.1-3 abind_1.4-8
[29] GenomicRanges_1.62.1 Rtsne_0.17
[31] BiocGenerics_0.56.0 circlize_0.4.16
[33] GenomeInfoDbData_1.2.14 IRanges_2.44.0
[35] S4Vectors_0.48.0 ggrepel_0.9.6
[37] irlba_2.3.5.1 listenv_0.10.0
[39] spatstat.utils_3.2-1 goftest_1.2-3
[41] RSpectra_0.16-2 spatstat.random_3.4-3
[43] fitdistrplus_1.2-2 parallelly_1.46.0
[45] codetools_0.2-20 DelayedArray_0.36.0
[47] tidyselect_1.2.1 shape_1.4.6.1
[49] UCSC.utils_1.4.0 farver_2.1.2
[51] viridis_0.6.5 ScaledMatrix_1.16.0
[53] matrixStats_1.5.0 stats4_4.5.2
[55] spatstat.explore_3.7-0 Seqinfo_1.0.0
[57] jsonlite_2.0.0 GetoptLong_1.1.0
[59] progressr_0.18.0 ggridges_0.5.6
[61] survival_3.8-3 iterators_1.0.14
[63] foreach_1.5.2 tools_4.5.2
[65] ica_1.0-3 Rcpp_1.1.0
[67] glue_1.8.0 gridExtra_2.3
[69] SparseArray_1.10.8 xfun_0.52
[71] distributional_0.6.0 MatrixGenerics_1.22.0
[73] ggthemes_5.2.0 GenomeInfoDb_1.44.0
[75] withr_3.0.2 fastmap_1.2.0
[77] digest_0.6.37 rsvd_1.0.5
[79] parallelDist_0.2.6 timechange_0.3.0
[81] R6_2.6.1 mime_0.13
[83] colorspace_2.1-1 Cairo_1.7-0
[85] scattermore_1.2 tensor_1.5
[87] spatstat.data_3.1-9 generics_0.1.3
[89] data.table_1.18.0 httr_1.4.7
[91] htmlwidgets_1.6.4 S4Arrays_1.10.1
[93] uwot_0.2.3 pkgconfig_2.0.3
[95] gtable_0.3.6 lmtest_0.9-40
[97] S7_0.2.1 SingleCellExperiment_1.32.0
[99] XVector_0.50.0 htmltools_0.5.8.1
[101] dotCall64_1.2 zigg_0.0.2
[103] clue_0.3-66 Biobase_2.70.0
[105] SeuratObject_5.3.0 scales_1.4.0
[107] png_0.1-8 spatstat.univar_3.1-6
[109] knitr_1.50 tzdb_0.5.0
[111] reshape2_1.4.4 rjson_0.2.23
[113] nlme_3.1-168 proxy_0.4-29
[115] cachem_1.1.0 zoo_1.8-14
[117] GlobalOptions_0.1.2 KernSmooth_2.23-26
[119] parallel_4.5.2 miniUI_0.1.2
[121] pillar_1.11.1 vctrs_0.7.0
[123] RANN_2.6.2 promises_1.5.0
[125] BiocSingular_1.26.1 beachmat_2.26.0
[127] xtable_1.8-4 cluster_2.1.8.1
[129] evaluate_1.0.5 magick_2.9.0
[131] cli_3.6.5 compiler_4.5.2
[133] rlang_1.1.7 crayon_1.5.3
[135] future.apply_1.20.0 plyr_1.8.9
[137] stringi_1.8.7 viridisLite_0.4.2
[139] deldir_2.0-4 BiocParallel_1.44.0
[141] lazyeval_0.2.2 spatstat.geom_3.7-0
[143] RcppHNSW_0.6.0 hms_1.1.3
[145] patchwork_1.3.2 future_1.69.0
[147] shiny_1.12.1 SummarizedExperiment_1.38.1
[149] ROCR_1.0-11 Rfast_2.1.5.1
[151] igraph_2.2.1 memoise_2.0.1
[153] RcppParallel_5.1.10